Repair of an Attenuated Low-Passage Murine Cytomegalovirus Bacterial Artificial Chromosome Identifies a Novel Spliced Gene Essential for Salivary Gland Tropism
- PMID: 32847854
- PMCID: PMC7592224
- DOI: 10.1128/JVI.01456-20
Repair of an Attenuated Low-Passage Murine Cytomegalovirus Bacterial Artificial Chromosome Identifies a Novel Spliced Gene Essential for Salivary Gland Tropism
Abstract
The cloning of herpesviruses as bacterial artificial chromosomes (BACs) has revolutionized the study of herpesvirus biology, allowing rapid and precise manipulation of viral genomes. Several clinical strains of human cytomegalovirus (HCMV) have been cloned as BACs; however, no low-passage strains of murine CMV (MCMV), which provide a model mimicking these isolates, have been cloned. Here, the low-passage G4 strain of was BAC cloned. G4 carries an m157 gene that does not ligate the natural killer (NK) cell-activating receptor, Ly49H, meaning that unlike laboratory strains of MCMV, this virus replicates well in C57BL/6 mice. This BAC clone exhibited normal replication during acute infection in the spleen and liver but was attenuated for salivary gland tropism. Next-generation sequencing revealed a C-to-A mutation at nucleotide position 188422, located in the 3' untranslated region of sgg1, a spliced gene critical for salivary gland tropism. Repair of this mutation restored tropism for the salivary glands. Transcriptional analysis revealed a novel spliced gene within the sgg1 locus. This small open reading frame (ORF), sgg1.1, starts at the 3' end of the first exon of sgg1 and extends exon 2 of sgg1. This shorter spliced gene is prematurely terminated by the nonsense mutation at nt 188422. Sequence analysis of tissue culture-passaged virus demonstrated that sgg1.1 was stable, although other mutational hot spots were identified. The G4 BAC will allow in vivo studies in a broader range of mice, avoiding the strong NK cell responses seen in B6 mice with other MCMV BAC-derived MCMVs.IMPORTANCE Murine cytomegalovirus (MCMV) is widely used as a model of human CMV (HCMV) infection. However, this model relies on strains of MCMV that have been serially passaged in the laboratory for over four decades. These laboratory strains have been cloned as bacterial artificial chromosomes (BACs), which permits rapid and precise manipulation. Low-passage strains of MCMV add to the utility of the mouse model of HCMV infection but do not exist as cloned BACs. This study describes the first such low-passage MCMV BAC. This BAC-derived G4 was initially attenuated in vivo, with subsequent full genomic sequencing revealing a novel spliced transcript required for salivary gland tropism. These data suggest that MCMV, like HCMV, undergoes tissue culture adaptation that can limit in vivo growth and supports the use of BAC clones as a way of standardizing viral strains and minimizing interlaboratory strain variation.
Keywords: BAC; G4; MCMV; sgg1; sgg1.1; tissue tropism.
Copyright © 2020 American Society for Microbiology.
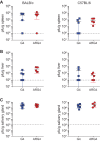
Figures



Similar articles
-
Mouse cytomegalovirus lacking sgg1 shows reduced import into the salivary glands.J Gen Virol. 2024 Aug;105(8):002013. doi: 10.1099/jgv.0.002013. J Gen Virol. 2024. PMID: 39093048 Free PMC article.
-
Virus progeny of murine cytomegalovirus bacterial artificial chromosome pSM3fr show reduced growth in salivary Glands due to a fixed mutation of MCK-2.J Virol. 2011 Oct;85(19):10346-53. doi: 10.1128/JVI.00545-11. Epub 2011 Aug 3. J Virol. 2011. PMID: 21813614 Free PMC article.
-
Structure and function of the murine cytomegalovirus sgg1 gene: a determinant of viral growth in salivary gland acinar cells.J Virol. 1994 Dec;68(12):7717-27. doi: 10.1128/JVI.68.12.7717-7727.1994. J Virol. 1994. PMID: 7966561 Free PMC article.
-
Human cytomegalovirus: taking the strain.Med Microbiol Immunol. 2015 Jun;204(3):273-84. doi: 10.1007/s00430-015-0411-4. Epub 2015 Apr 17. Med Microbiol Immunol. 2015. PMID: 25894764 Free PMC article. Review.
-
Conditional gene expression systems to study herpesvirus biology in vivo.Med Microbiol Immunol. 2008 Jun;197(2):269-76. doi: 10.1007/s00430-008-0086-1. Epub 2008 Mar 7. Med Microbiol Immunol. 2008. PMID: 18324415 Review.
Cited by
-
Mouse cytomegalovirus lacking sgg1 shows reduced import into the salivary glands.J Gen Virol. 2024 Aug;105(8):002013. doi: 10.1099/jgv.0.002013. J Gen Virol. 2024. PMID: 39093048 Free PMC article.
-
Characterization of M116.1p, a Murine Cytomegalovirus Protein Required for Efficient Infection of Mononuclear Phagocytes.J Virol. 2022 Jan 26;96(2):e0087621. doi: 10.1128/JVI.00876-21. Epub 2021 Oct 27. J Virol. 2022. PMID: 34705561 Free PMC article.
-
A Review of Murine Cytomegalovirus as a Model for Human Cytomegalovirus Disease-Do Mice Lie?Int J Mol Sci. 2020 Dec 28;22(1):214. doi: 10.3390/ijms22010214. Int J Mol Sci. 2020. PMID: 33379272 Free PMC article. Review.
-
Immune surveillance of cytomegalovirus in tissues.Cell Mol Immunol. 2024 Sep;21(9):959-981. doi: 10.1038/s41423-024-01186-2. Epub 2024 Aug 12. Cell Mol Immunol. 2024. PMID: 39134803 Free PMC article. Review.
-
Fine-tuning the evolutionary stability of recombinant herpesviral transmissible vaccines.Proc Biol Sci. 2024 Nov;291(2034):20241827. doi: 10.1098/rspb.2024.1827. Epub 2024 Nov 13. Proc Biol Sci. 2024. PMID: 39532136 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

