HPO iron chelator, CP655, causes the G1/S phase cell cycle block via p21 upregulation
- PMID: 32865890
- PMCID: PMC7654408
- DOI: 10.1002/iid3.342
HPO iron chelator, CP655, causes the G1/S phase cell cycle block via p21 upregulation
Abstract
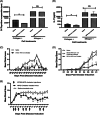
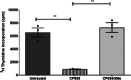
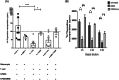
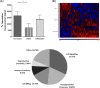
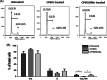
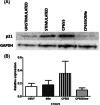
Iron is known not only for its importance in cellular and metabolic pathways but also for its role in causing cellular toxicities such as production of reactive oxygen species and growth of pathogens. The inability of the human body to physiologically excrete excess iron highlights the need to develop a cheap yet effective iron chelator. This study provides initial evidence of the therapeutic and prophylactic properties of 3-hydroxypyridin-4-one (HPO) chelators in murine collagen-induced arthritis. To determine whether these chelators would be effective on human cells, we tested a panel of different HPO chelators and identified 7-diethylamino-N-((5-hydroxy-6-methyl-4-oxo-1,4-dihydropyridin-3-yl)methyl)-N-methyl-2-oxo-chromen-3-carboxamide (CP655) as the most effective compound targeting human CD4+ T cells. Treatment with CP655 causes significant inhibition of cell proliferation and production of inflammatory cytokines such as interferon-γ and interleukin-17. Microarray analysis revealed dysregulation in cell cycle-related genes following CP655 treatment. This was validated by flow cytometry demonstrating a G1/S phase block caused by CP655. Finally, mechanistic experiments revealed that the chelator may be causing an upregulation of the cell cycle inhibitor protein CDKN1A (p21) as a possible mechanism of action. In conclusion, this study demonstrates that HPO chelators could prove to have therapeutic potential for diseases driven by excessive T cell proliferation.
Keywords: CP655; HPO chelators; T cell proliferation; cell cycle; iron chelation; iron metabolism; p21.
© 2020 The Authors. Immunity, Inflammation and Disease published by John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare that there are no conflict of interests.
Figures

References
-
- Puig S, Ramos‐Alonso L, Romero AM, Martínez‐Pastor MT. The elemental role of iron in DNA synthesis and repair. Metallomics. 2017;9(11):1483‐1500. - PubMed
-
- Cao LL, Liu H, Yue Z, et al. Iron chelation inhibits cancer cell growth and modulates global histone methylation status in colorectal cancer. BioMetals. 2018;31(5):797‐805. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

