Structure of the Capsid Size-Determining Scaffold of "Satellite" Bacteriophage P4
- PMID: 32867300
- PMCID: PMC7552001
- DOI: 10.3390/v12090953
Structure of the Capsid Size-Determining Scaffold of "Satellite" Bacteriophage P4
Abstract
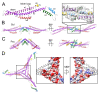
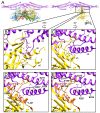
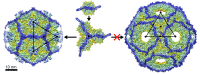
P4 is a mobile genetic element (MGE) that can exist as a plasmid or integrated into its Escherichia coli host genome, but becomes packaged into phage particles by a helper bacteriophage, such as P2. P4 is the original example of what we have termed "molecular piracy", the process by which one MGE usurps the life cycle of another for its own propagation. The P2 helper provides most of the structural gene products for assembly of the P4 virion. However, when P4 is mobilized by P2, the resulting capsids are smaller than those normally formed by P2 alone. The P4-encoded protein responsible for this size change is called Sid, which forms an external scaffolding cage around the P4 procapsids. We have determined the high-resolution structure of P4 procapsids, allowing us to build an atomic model for Sid as well as the gpN capsid protein. Sixty copies of Sid form an intertwined dodecahedral cage around the T = 4 procapsid, making contact with only one out of the four symmetrically non-equivalent copies of gpN. Our structure provides a basis for understanding the sir mutants in gpN that prevent small capsid formation, as well as the nms "super-sid" mutations that counteract the effect of the sir mutations, and suggests a model for capsid size redirection by Sid.
Keywords: Caudovirales; Psu; Sid; bacteriophage P2; capsid assembly; mobile genetic elements; molecular piracy; sir mutants; size determination.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



Similar articles
-
The structure of P4 procapsids produced by coexpression of capsid and external scaffolding proteins.Virology. 2002 Jul 5;298(2):224-31. doi: 10.1006/viro.2002.1485. Virology. 2002. PMID: 12127785
-
In vitro assembly of bacteriophage P4 procapsids from purified capsid and scaffolding proteins.Virology. 2000 Sep 15;275(1):133-44. doi: 10.1006/viro.2000.0521. Virology. 2000. PMID: 11017795
-
Structure and size determination of bacteriophage P2 and P4 procapsids: function of size responsiveness mutations.J Struct Biol. 2012 Jun;178(3):215-24. doi: 10.1016/j.jsb.2012.04.002. Epub 2012 Apr 9. J Struct Biol. 2012. PMID: 22508104 Free PMC article.
-
Mechanisms of genome propagation and helper exploitation by satellite phage P4.Microbiol Rev. 1993 Sep;57(3):683-702. doi: 10.1128/mr.57.3.683-702.1993. Microbiol Rev. 1993. PMID: 8246844 Free PMC article. Review.
-
Pirates of the Caudovirales.Virology. 2012 Dec 20;434(2):210-21. doi: 10.1016/j.virol.2012.10.028. Epub 2012 Nov 3. Virology. 2012. PMID: 23131350 Free PMC article. Review.
Cited by
-
Phage-mediated horizontal gene transfer and its implications for the human gut microbiome.Gastroenterol Rep (Oxf). 2022 Apr 13;10:goac012. doi: 10.1093/gastro/goac012. eCollection 2022. Gastroenterol Rep (Oxf). 2022. PMID: 35425613 Free PMC article.
-
A Vibrio cholerae viral satellite maximizes its spread and inhibits phage by remodeling hijacked phage coat proteins into small capsids.Elife. 2024 Jan 11;12:RP87611. doi: 10.7554/eLife.87611. Elife. 2024. PMID: 38206122 Free PMC article.
-
High Resolution Structure of the Mature Capsid of Ralstonia solanacearum Bacteriophage ϕRSA1 by Cryo-Electron Microscopy.Int J Mol Sci. 2021 Oct 13;22(20):11053. doi: 10.3390/ijms222011053. Int J Mol Sci. 2021. PMID: 34681713 Free PMC article.
-
Elucidating double stranded DNA viral scaffolding protein structures through advances in cryogenic electron microscopy data processing.Curr Opin Struct Biol. 2025 Jun 16;94:103081. doi: 10.1016/j.sbi.2025.103081. Online ahead of print. Curr Opin Struct Biol. 2025. PMID: 40527155 Review.
-
Shape shifter: redirection of prolate phage capsid assembly by staphylococcal pathogenicity islands.Nat Commun. 2021 Nov 4;12(1):6408. doi: 10.1038/s41467-021-26759-x. Nat Commun. 2021. PMID: 34737316 Free PMC article.
References
-
- Fane B.A., Brentlinger K.L., Burch A.D., Chen M., Hafenstein S., Moore E., Novak C.R., Uchiyama A. The Microviridae. In: Calendar R., editor. The Bacteriophages. Oxford University Press; New York, NY, USA: 2006. pp. 129–145.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

