Organic Solvents for Enhanced Proteolysis of Stable Proteins for Hydrogen-Deuterium Exchange Mass Spectrometry
- PMID: 32867496
- PMCID: PMC7485609
- DOI: 10.1021/acs.analchem.0c02194
Organic Solvents for Enhanced Proteolysis of Stable Proteins for Hydrogen-Deuterium Exchange Mass Spectrometry
Abstract
Protein digestion is a key challenge in mass spectrometry (MS)-based structural proteomics. Although using hydrogen-deuterium exchange kinetics with MS (HDX-MS) to interrogate the high-order structure of proteins is now established, it can be challenging for β-barrel proteins, which are important in cellular transport. These proteins contain a continuous chain of H-bonds that impart stability, causing difficulty in digestion for bottom-up measurements. To overcome this impediment, we tested organic solvents as denaturants during on-line pepsin digestion of soluble β-barrel proteins. We selected green fluorescent protein (GFP), siderocalin (Scn), and retinol-binding protein 4 (RBP4) as model proteins and screened six different polar-aprotic and polar-protic solvent combinations to disrupt the H-bonds and hydrophobic interactions holding together the β-sheets. The use of organic solvents improves digestion, generating more peptides from the rigid β-barrel regions, without compromising the ability to predict the retinol binding site on RBP4 when adopting this proteolysis with HDX.
Figures
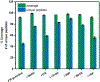
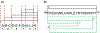
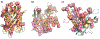

References
-
- Percya A; Reyb M; Burnsb K; Schriemer D Probing protein interactions with hydrogen/deuterium exchange and mass spectrometry—A review. Anal. Chim. Acta 2012, 721, 6, 7–21. - PubMed
-
- Giansanti P; Tsiatsiani L; Low T; Heck A Six alternative proteases for mass spectrometry–based proteomics beyond trypsin. Nat. Protoc 2016, 11, 993–1006. - PubMed
-
- Serpa J; Parker C; Petrotchenko E; Han J; Pan J; Borchersa C Mass spectrometry-based structural proteomics. Eur. J. Mass Spectrom 2012, 18, 251–267. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

