eQTL Colocalization Analyses Identify NTN4 as a Candidate Breast Cancer Risk Gene
- PMID: 32871102
- PMCID: PMC7536644
- DOI: 10.1016/j.ajhg.2020.08.006
eQTL Colocalization Analyses Identify NTN4 as a Candidate Breast Cancer Risk Gene
Abstract
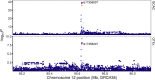
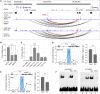
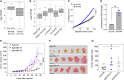
Breast cancer genome-wide association studies (GWASs) have identified 150 genomic risk regions containing more than 13,000 credible causal variants (CCVs). The CCVs are predominantly noncoding and enriched in regulatory elements. However, the genes underlying breast cancer risk associations are largely unknown. Here, we used genetic colocalization analysis to identify loci at which gene expression could potentially explain breast cancer risk phenotypes. Using data from the Breast Cancer Association Consortium (BCAC) and quantitative trait loci (QTL) from the Genotype-Tissue Expression (GTEx) project and The Cancer Genome Project (TCGA), we identify shared genetic relationships and reveal novel associations between cancer phenotypes and effector genes. Seventeen genes, including NTN4, were identified as potential mediators of breast cancer risk. For NTN4, we showed the rs61938093 CCV at this region was located within an enhancer element that physically interacts with the NTN4 promoter, and the risk allele reduced NTN4 promoter activity. Furthermore, knockdown of NTN4 in breast cells increased cell proliferation in vitro and tumor growth in vivo. These data provide evidence linking risk-associated variation to genes that may contribute to breast cancer predisposition.
Keywords: GWAS; NTN4; breast cancer; colocalization; eQTL; enhancer; tumor suppressor.
Crown Copyright © 2020. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
The authors declare no competing interests.
Figures

Similar articles
-
The risk variant rs11836367 contributes to breast cancer onset and metastasis by attenuating Wnt signaling via regulating NTN4 expression.Sci Adv. 2022 Jun 10;8(23):eabn3509. doi: 10.1126/sciadv.abn3509. Epub 2022 Jun 10. Sci Adv. 2022. PMID: 35687692 Free PMC article.
-
A Comprehensive cis-eQTL Analysis Revealed Target Genes in Breast Cancer Susceptibility Loci Identified in Genome-wide Association Studies.Am J Hum Genet. 2018 May 3;102(5):890-903. doi: 10.1016/j.ajhg.2018.03.016. Am J Hum Genet. 2018. PMID: 29727689 Free PMC article.
-
Risk SNP-Mediated Enhancer-Promoter Interaction Drives Colorectal Cancer through Both FADS2 and AP002754.2.Cancer Res. 2020 May 1;80(9):1804-1818. doi: 10.1158/0008-5472.CAN-19-2389. Epub 2020 Mar 3. Cancer Res. 2020. PMID: 32127356
-
Integrative eQTL-based analyses reveal the biology of breast cancer risk loci.Cell. 2013 Jan 31;152(3):633-41. doi: 10.1016/j.cell.2012.12.034. Cell. 2013. PMID: 23374354 Free PMC article. Review.
-
Inherited genetic susceptibility to breast cancer: the beginning of the end or the end of the beginning?Am J Pathol. 2013 Oct;183(4):1038-1051. doi: 10.1016/j.ajpath.2013.07.003. Epub 2013 Aug 23. Am J Pathol. 2013. PMID: 23973388 Review.
Cited by
-
METRO: Multi-ancestry transcriptome-wide association studies for powerful gene-trait association detection.Am J Hum Genet. 2022 May 5;109(5):783-801. doi: 10.1016/j.ajhg.2022.03.003. Epub 2022 Mar 24. Am J Hum Genet. 2022. PMID: 35334221 Free PMC article.
-
A genome-wide association study of mammographic texture variation.Breast Cancer Res. 2022 Nov 7;24(1):76. doi: 10.1186/s13058-022-01570-8. Breast Cancer Res. 2022. PMID: 36344993 Free PMC article.
-
Identification of target proteins for breast cancer genetic risk loci and blood risk biomarkers in a large study by integrating genomic and proteomic data.Int J Cancer. 2023 Jun 1;152(11):2314-2320. doi: 10.1002/ijc.34472. Epub 2023 Feb 21. Int J Cancer. 2023. PMID: 36779764 Free PMC article.
-
Netrin Family Genes as Prognostic Markers and Therapeutic Targets for Clear Cell Renal Cell Carcinoma: Netrin-4 Acts through the Wnt/β-Catenin Signaling Pathway.Cancers (Basel). 2023 May 18;15(10):2816. doi: 10.3390/cancers15102816. Cancers (Basel). 2023. PMID: 37345154 Free PMC article.
-
CRISPR screens identify gene targets at breast cancer risk loci.Genome Biol. 2023 Mar 29;24(1):59. doi: 10.1186/s13059-023-02898-w. Genome Biol. 2023. PMID: 36991492 Free PMC article.
References
-
- Ongen H., Brown A.A., Delaneau O., Panousis N.I., Nica A.C., Dermitzakis E.T., GTEx Consortium Estimating the causal tissues for complex traits and diseases. Nat. Genet. 2017;49:1676–1683. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

