DynaMut2: Assessing changes in stability and flexibility upon single and multiple point missense mutations
- PMID: 32881105
- PMCID: PMC7737773
- DOI: 10.1002/pro.3942
DynaMut2: Assessing changes in stability and flexibility upon single and multiple point missense mutations
Abstract
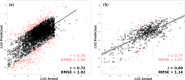
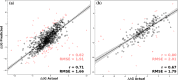
Predicting the effect of missense variations on protein stability and dynamics is important for understanding their role in diseases, and the link between protein structure and function. Approaches to estimate these changes have been proposed, but most only consider single-point missense variants and a static state of the protein, with those that incorporate dynamics are computationally expensive. Here we present DynaMut2, a web server that combines Normal Mode Analysis (NMA) methods to capture protein motion and our graph-based signatures to represent the wildtype environment to investigate the effects of single and multiple point mutations on protein stability and dynamics. DynaMut2 was able to accurately predict the effects of missense mutations on protein stability, achieving Pearson's correlation of up to 0.72 (RMSE: 1.02 kcal/mol) on a single point and 0.64 (RMSE: 1.80 kcal/mol) on multiple-point missense mutations across 10-fold cross-validation and independent blind tests. For single-point mutations, DynaMut2 achieved comparable performance with other methods when predicting variations in Gibbs Free Energy (ΔΔG) and in melting temperature (ΔTm ). We anticipate our tool to be a valuable suite for the study of protein flexibility analysis and the study of the role of variants in disease. DynaMut2 is freely available as a web server and API at http://biosig.unimelb.edu.au/dynamut2.
Keywords: dynamics; graph-based signatures; missense mutations; stability changes.
© 2020 The Authors. Protein Science published by Wiley Periodicals LLC on behalf of The Protein Society.
Conflict of interest statement
No conflict of interest declared.
Figures

Similar articles
-
mmCSM-PPI: predicting the effects of multiple point mutations on protein-protein interactions.Nucleic Acids Res. 2021 Jul 2;49(W1):W417-W424. doi: 10.1093/nar/gkab273. Nucleic Acids Res. 2021. PMID: 33893812 Free PMC article.
-
DynaMut: predicting the impact of mutations on protein conformation, flexibility and stability.Nucleic Acids Res. 2018 Jul 2;46(W1):W350-W355. doi: 10.1093/nar/gky300. Nucleic Acids Res. 2018. PMID: 29718330 Free PMC article.
-
DDMut: predicting effects of mutations on protein stability using deep learning.Nucleic Acids Res. 2023 Jul 5;51(W1):W122-W128. doi: 10.1093/nar/gkad472. Nucleic Acids Res. 2023. PMID: 37283042 Free PMC article.
-
Molecular mechanisms of disease-causing missense mutations.J Mol Biol. 2013 Nov 1;425(21):3919-36. doi: 10.1016/j.jmb.2013.07.014. Epub 2013 Jul 16. J Mol Biol. 2013. PMID: 23871686 Free PMC article. Review.
-
Computational Approaches to Prioritize Cancer Driver Missense Mutations.Int J Mol Sci. 2018 Jul 20;19(7):2113. doi: 10.3390/ijms19072113. Int J Mol Sci. 2018. PMID: 30037003 Free PMC article. Review.
Cited by
-
FireProt 2.0: web-based platform for the fully automated design of thermostable proteins.Brief Bioinform. 2023 Nov 22;25(1):bbad425. doi: 10.1093/bib/bbad425. Brief Bioinform. 2023. PMID: 38018911 Free PMC article.
-
Infection by a multidrug-resistant Corynebacterium diphtheriae strain: prediction of virulence factors, CRISPR-Cas system analysis, and structural implications of mutations conferring rifampin resistance.Funct Integr Genomics. 2024 Aug 28;24(5):145. doi: 10.1007/s10142-024-01434-8. Funct Integr Genomics. 2024. PMID: 39196424
-
Analysis of proteins in the light of mutations.Eur Biophys J. 2024 Aug;53(5-6):255-265. doi: 10.1007/s00249-024-01714-y. Epub 2024 Jul 2. Eur Biophys J. 2024. PMID: 38955858 Review.
-
Identification and clinical characteristics of a novel missense ADGRG1 variant in bilateral Frontoparietal Polymicrogyria: The electroclinical change from infancy to adulthood after Callosotomy in three siblings.Epilepsia Open. 2023 Mar;8(1):154-164. doi: 10.1002/epi4.12685. Epub 2023 Jan 11. Epilepsia Open. 2023. PMID: 36524291 Free PMC article. Review.
-
Characterizing and explaining the impact of disease-associated mutations in proteins without known structures or structural homologs.Brief Bioinform. 2022 Jul 18;23(4):bbac187. doi: 10.1093/bib/bbac187. Brief Bioinform. 2022. PMID: 35641150 Free PMC article.
References
-
- Byrne JA, Strautnieks SS, Ihrke G, et al. Missense mutations and single nucleotide polymorphisms in ABCB11 impair bile salt export pump processing and function or disrupt pre‐messenger RNA splicing. Hepatology. 2009;49:553–567. - PubMed
-
- Jafri M, Wake NC, Ascher DB, et al. Germline mutations in the CDKN2B tumor suppressor gene predispose to renal cell carcinoma. Cancer Discov. 2015;5:723–729. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

