TBDB: a database of structurally annotated T-box riboswitch:tRNA pairs
- PMID: 32882008
- PMCID: PMC7778990
- DOI: 10.1093/nar/gkaa721
TBDB: a database of structurally annotated T-box riboswitch:tRNA pairs
Abstract
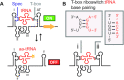
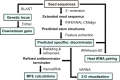
T-box riboswitches constitute a large family of tRNA-binding leader sequences that play a central role in gene regulation in many gram-positive bacteria. Accurate inference of the tRNA binding to T-box riboswitches is critical to predict their cis-regulatory activity. However, there is no central repository of information on the tRNA binding specificities of T-box riboswitches, and de novo prediction of binding specificities requires advanced knowledge of computational tools to annotate riboswitch secondary structure features. Here, we present the T-box Riboswitch Annotation Database (TBDB, https://tbdb.io), an open-access database with a collection of 23,535 T-box riboswitch sequences, spanning the major phyla of 3,632 bacterial species. Among structural predictions, the TBDB also identifies specifier sequences, cognate tRNA binding partners, and downstream regulatory targets. To our knowledge, the TBDB presents the largest collection of feature, sequence, and structural annotations carried out on this important family of regulatory RNA.
© The Author(s) 2020. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures
References
-
- Winkler W. Riboswitches and the role of noncoding RNAs in bacterial metabolic control. Curr. Opin. Chem. Biol. 2005; 9:594–602. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

