Limited proliferation capacity of aortic intima resident macrophages requires monocyte recruitment for atherosclerotic plaque progression
- PMID: 32895539
- PMCID: PMC7502558
- DOI: 10.1038/s41590-020-0768-4
Limited proliferation capacity of aortic intima resident macrophages requires monocyte recruitment for atherosclerotic plaque progression
Abstract
Early atherosclerosis depends upon responses by immune cells resident in the intimal aortic wall. Specifically, the healthy intima is thought to be populated by vascular dendritic cells (DCs) that, during hypercholesterolemia, initiate atherosclerosis by being the first to accumulate cholesterol. Whether these cells remain key players in later stages of disease is unknown. Using murine lineage-tracing models and gene expression profiling, we reveal that myeloid cells present in the intima of the aortic arch are not DCs but instead specialized aortic intima resident macrophages (MacAIR) that depend upon colony-stimulating factor 1 and are sustained by local proliferation. Although MacAIR comprise the earliest foam cells in plaques, their proliferation during plaque progression is limited. After months of hypercholesterolemia, their presence in plaques is overtaken by recruited monocytes, which induce MacAIR-defining genes. These data redefine the lineage of intimal phagocytes and suggest that proliferation is insufficient to sustain generations of macrophages during plaque progression.
Figures



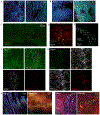

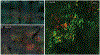



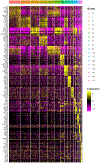




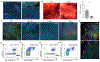


References
-
- Geovanini GR & Libby P Atherosclerosis and inflammation: Overview and updates. Clinical Science 132, 1243–1252 (2018). - PubMed
-
- Benjamin EJ et al. Heart Disease and Stroke Statistics-2019 Update: A Report From the American Heart Association. Circulation 139, e56–e66 (2019). - PubMed
-
- Ridker PM et al. Antiinflammatory therapy with canakinumab for atherosclerotic disease. N. Engl. J. Med 377, 1119–1131 (2017). - PubMed
Online References
-
- Mackarehtschian K et al. Targeted disruption of the flk2/flt3 gene leads to deficiencies in primitive hematopoietic progenitors. Immunity 3, 147–161 (1995). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- 17POST33410473/AHA/American Heart Association-American Stroke Association/United States
- R37 AI049653/AI/NIAID NIH HHS/United States
- R00 HL138163/HL/NHLBI NIH HHS/United States
- T32 AI007313/AI/NIAID NIH HHS/United States
- K99 HL138163/HL/NHLBI NIH HHS/United States
- K08 HL123519/HL/NHLBI NIH HHS/United States
- R01 HL139714/HL/NHLBI NIH HHS/United States
- T32 DK007120/DK/NIDDK NIH HHS/United States
- P01 AI035296/AI/NIAID NIH HHS/United States
- DP1 DK109668/DK/NIDDK NIH HHS/United States
- R01 HL138466/HL/NHLBI NIH HHS/United States
- P30 DK020579/DK/NIDDK NIH HHS/United States
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

