Autophagy involvement in oncogenesis
- PMID: 32897597
- PMCID: PMC7648016
- DOI: 10.1111/cas.14646
Autophagy involvement in oncogenesis
Abstract
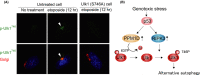
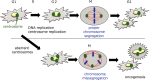
Various clinical and experimental findings have revealed the causal relationship between autophagy failure and oncogenesis, and several mechanisms have been suggested to explain this relationship. We recently proposed two additional mechanisms: centrosome number dysregulation and the failure of autophagic cell death. Here, we detail the mechanical relationship between autophagy failure and oncogenesis.
Keywords: alternative autophagy; autophagic cell death; autophagy; centrosome; oncogenesis.
© 2020 The Authors. Cancer Science published by John Wiley & Sons Australia, Ltd on behalf of Japanese Cancer Association.
Conflict of interest statement
The authors declare no conflicts of interest associated with this manuscript.
Figures



References
-
- Ohsumi Y. Molecular dissection of autophagy: two ubiquitin‐like systems. Nat Rev Mol Cell Biol. 2001;2:211‐216. - PubMed
-
- Mizushima N, Yoshimori T, Ohsumi Y. The role of Atg proteins in autophagosome formation. Annu Rev Cell Dev Biol. 2011;27:107‐132. - PubMed
-
- Shimizu S, Kanaseki T, Mizushima N, et al. Role of Bcl‐2 family proteins in a non‐apoptotic programmed cell death dependent on autophagy genes. Nat Cell Biol. 2004;6:1221‐1228. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

