In vivo antiviral host transcriptional response to SARS-CoV-2 by viral load, sex, and age
- PMID: 32898168
- PMCID: PMC7478592
- DOI: 10.1371/journal.pbio.3000849
In vivo antiviral host transcriptional response to SARS-CoV-2 by viral load, sex, and age
Abstract
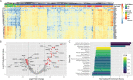
Despite limited genomic diversity, severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has shown a wide range of clinical manifestations in different patient populations. The mechanisms behind these host differences are still unclear. Here, we examined host response gene expression across infection status, viral load, age, and sex among shotgun RNA sequencing profiles of nasopharyngeal (NP) swabs from 430 individuals with PCR-confirmed SARS-CoV-2 and 54 negative controls. SARS-CoV-2 induced a strong antiviral response with up-regulation of antiviral factors such as OAS1-3 and IFIT1-3 and T helper type 1 (Th1) chemokines CXCL9/10/11, as well as a reduction in transcription of ribosomal proteins. SARS-CoV-2 culture in human airway epithelial (HAE) cultures replicated the in vivo antiviral host response 7 days post infection, with no induction of interferon-stimulated genes after 3 days. Patient-matched longitudinal specimens (mean elapsed time = 6.3 days) demonstrated reduction in interferon-induced transcription, recovery of transcription of ribosomal proteins, and initiation of wound healing and humoral immune responses. Expression of interferon-responsive genes, including ACE2, increased as a function of viral load, while transcripts for B cell-specific proteins and neutrophil chemokines were elevated in patients with lower viral load. Older individuals had reduced expression of the Th1 chemokines CXCL9/10/11 and their cognate receptor CXCR3, as well as CD8A and granzyme B, suggesting deficiencies in trafficking and/or function of cytotoxic T cells and natural killer (NK) cells. Relative to females, males had reduced B cell-specific and NK cell-specific transcripts and an increase in inhibitors of nuclear factor kappa-B (NF-κB) signaling, possibly inappropriately throttling antiviral responses. Collectively, our data demonstrate that host responses to SARS-CoV-2 are dependent on viral load and infection time course, with observed differences due to age and sex that may contribute to disease severity.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures




Update of
-
In vivo antiviral host response to SARS-CoV-2 by viral load, sex, and age.bioRxiv [Preprint]. 2020 Jun 22:2020.06.22.165225. doi: 10.1101/2020.06.22.165225. bioRxiv. 2020. Update in: PLoS Biol. 2020 Sep 8;18(9):e3000849. doi: 10.1371/journal.pbio.3000849. PMID: 32607510 Free PMC article. Updated. Preprint.
References
-
- OpenSAFELY: factors associated with COVID-19-related hospital death in the linked electronic health records of 17 million adult NHS patients. | medRxiv [Internet]. [cited 2020 Jun 5]. Available from: https://www.medrxiv.org/content/10.1101/2020.05.06.20092999v1. - DOI
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

