Targeting Nonsense: Optimization of 1,2,4-Oxadiazole TRIDs to Rescue CFTR Expression and Functionality in Cystic Fibrosis Cell Model Systems
- PMID: 32899265
- PMCID: PMC7504161
- DOI: 10.3390/ijms21176420
Targeting Nonsense: Optimization of 1,2,4-Oxadiazole TRIDs to Rescue CFTR Expression and Functionality in Cystic Fibrosis Cell Model Systems
Abstract
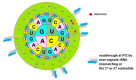
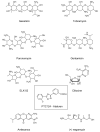
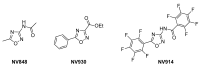
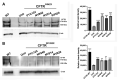
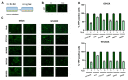
Cystic fibrosis (CF) patients develop a severe form of the disease when the cystic fibrosis transmembrane conductance regulator (CFTR) gene is affected by nonsense mutations. Nonsense mutations are responsible for the presence of a premature termination codon (PTC) in the mRNA, creating a lack of functional protein. In this context, translational readthrough-inducing drugs (TRIDs) represent a promising approach to correct the basic defect caused by PTCs. By using computational optimization and biological screening, we identified three new small molecules showing high readthrough activity. The activity of these compounds has been verified by evaluating CFTR expression and functionality after treatment with the selected molecules in cells expressing nonsense-CFTR-mRNA. Additionally, the channel functionality was measured by the halide sensitive yellow fluorescent protein (YFP) quenching assay. All three of the new TRIDs displayed high readthrough activity and low toxicity and can be considered for further evaluation as a therapeutic approach toward the second major cause of CF.
Keywords: genetic disorder; nonsense mutation; oxadiazoles; premature termination codon; translational readthrough inducing drugs.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

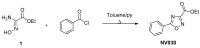
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

