Differential regulation of rho GTPases during lung adenocarcinoma migration and invasion reveals a novel role of the tumor suppressor StarD13 in invadopodia regulation
- PMID: 32900380
- PMCID: PMC7487901
- DOI: 10.1186/s12964-020-00635-5
Differential regulation of rho GTPases during lung adenocarcinoma migration and invasion reveals a novel role of the tumor suppressor StarD13 in invadopodia regulation
Abstract
Background: Lung cancer is the second most commonly occurring cancer. The ability to metastasize and spread to distant locations renders the tumor more aggressive. Members of the Rho subfamily of small GTP-binding proteins (GTPases) play a central role in the regulation of the actin cytoskeleton and in cancer cell migration and metastasis. In this study we investigated the role of the RhoA/Cdc42 GAP, StarD13, a previously described tumor suppressor, in malignancy, migration and invasion of the lung cancer cells A549.
Methods: We knocked down StarD13 expression in A549 lung cancer cells and tested the effect on cell migration and invadopodia formation using time lapse imaging and invasion assays. We also performed rescue experiments to determine the signaling pathways downstream of StarD13 and transfected the cells with FRET biosensors for RhoGTPases to identify the proteins involved in invadopodia formation.
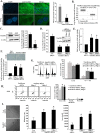
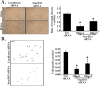
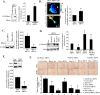
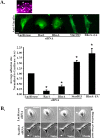
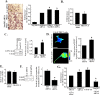
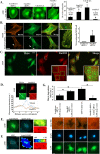
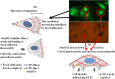
Results: We observed a decrease in the level of expression of StarD13 in lung tumor tissues compared to normal lung tissues through immunohistochemistry. StarD13 also showed a lower expression in the lung adenocarcinoma cell line A549 compared to normal lung cells, WI38. In addition, the depletion of StarD13 increased cell proliferation and viability in WI38 and A549 cells, suggesting that StarD13 might potentially be a tumor suppressor in lung cancer. The depletion of StarD13, however, inhibited cell motility, conversely demonstrating a positive regulatory role in cell migration. This was potentially due to the constitutive activation of RhoA detected by pull down and FRET assays. Surprisingly, StarD13 suppressed cell invasion by inhibiting Cdc42-mediated invadopodia formation. Indeed, TKS4 staining and invadopodia assay revealed that StarD13 depletion increased Cdc42 activation as well as invadopodia formation and matrix degradation. Normal lung cells depleted of StarD13 also produced invadopodia, otherwise a unique hallmark of invasive cancer cells. Cdc42 knock down mimicked the effects of StarD13, while overexpression of a constitutively active Cdc42 mimicked the effects of its depletion. Finally, immunostaining and FRET analysis revealed the absence of StarD13 in invadopodia as compared to Cdc42, which was activated in invadopodia at the sites of matrix degradation.
Conclusion: In conclusion, StarD13 plays distinct roles in lung cancer cell migration and invasion through its differential regulation of Rho GTPases. Video abstract.
Keywords: Invadopodia; Invasion; Migration; Rho GTPases; StarD13.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures
Similar articles
-
StarD13 negatively regulates invadopodia formation and invasion in high-grade serous (HGS) ovarian adenocarcinoma cells by inhibiting Cdc42.Eur J Cell Biol. 2022 Jan;101(1):151197. doi: 10.1016/j.ejcb.2021.151197. Epub 2021 Dec 21. Eur J Cell Biol. 2022. PMID: 34958986 Free PMC article.
-
StarD13 differentially regulates migration and invasion in prostate cancer cells.Hum Cell. 2021 Mar;34(2):607-623. doi: 10.1007/s13577-020-00479-8. Epub 2021 Jan 9. Hum Cell. 2021. PMID: 33420961
-
StarD13 is a tumor suppressor in breast cancer that regulates cell motility and invasion.Int J Oncol. 2014 May;44(5):1499-511. doi: 10.3892/ijo.2014.2330. Epub 2014 Mar 7. Int J Oncol. 2014. PMID: 24627003 Free PMC article.
-
StarD13: a potential star target for tumor therapeutics.Hum Cell. 2020 Jul;33(3):437-443. doi: 10.1007/s13577-020-00358-2. Epub 2020 Apr 9. Hum Cell. 2020. PMID: 32274657 Review.
-
The Role of Rho GTPases in Motility and Invasion of Glioblastoma Cells.Anal Cell Pathol (Amst). 2020 Jan 31;2020:9274016. doi: 10.1155/2020/9274016. eCollection 2020. Anal Cell Pathol (Amst). 2020. PMID: 32089990 Free PMC article. Review.
Cited by
-
Circ_0003570 Suppresses the progression of hepatocellular carcinoma through miR-182-5p/STARD13 regulatory axis.Biol Proced Online. 2022 Oct 14;24(1):14. doi: 10.1186/s12575-022-00176-w. Biol Proced Online. 2022. PMID: 36241975 Free PMC article.
-
ARHGAP17 regulates the spatiotemporal activity of Cdc42 at invadopodia.J Cell Biol. 2023 Feb 6;222(2):e202207020. doi: 10.1083/jcb.202207020. Epub 2022 Dec 26. J Cell Biol. 2023. PMID: 36571786 Free PMC article.
-
The 'Yin and Yang' of Cancer Cell Growth and Mechanosensing.Cancers (Basel). 2021 Sep 23;13(19):4754. doi: 10.3390/cancers13194754. Cancers (Basel). 2021. PMID: 34638240 Free PMC article. Review.
-
Inhibition of Cxcr4 Disrupts Mouse Embryonic Palatal Mesenchymal Cell Migration and Induces Cleft Palate Occurrence.Int J Mol Sci. 2023 Aug 13;24(16):12740. doi: 10.3390/ijms241612740. Int J Mol Sci. 2023. PMID: 37628919 Free PMC article.
-
StarD13 negatively regulates invadopodia formation and invasion in high-grade serous (HGS) ovarian adenocarcinoma cells by inhibiting Cdc42.Eur J Cell Biol. 2022 Jan;101(1):151197. doi: 10.1016/j.ejcb.2021.151197. Epub 2021 Dec 21. Eur J Cell Biol. 2022. PMID: 34958986 Free PMC article.
References
-
- Giese A, Bjerkvig R, Berens ME, Westphal M. Cost of migration: invasion of malignant gliomas and implications for treatment. J Clin Oncol. 2003;21(8):1624–1636. - PubMed
-
- Bailly M, Condeelis JS, Segall JE. Chemoattractant-induced lamellipod extension. Microsc Res Tech. 1998;43(5):433–443. - PubMed
-
- Gupton SL, Waterman-Storer CM. Spatiotemporal feedback between actomyosin and focal-adhesion systems optimizes rapid cell migration. Cell. 2006;125(7):1361–1374. - PubMed
-
- Condeelis JS, Wyckoff JB, Bailly M, Pestell R, Lawrence D, Backer J, et al. Lamellipodia in invasion. Semin Cancer Biol. 2001;11(2):119–128. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

