Ribonucleocapsid assembly/packaging signals in the genomes of the coronaviruses SARS-CoV and SARS-CoV-2: detection, comparison and implications for therapeutic targeting
- PMID: 32901577
- PMCID: PMC7544952
- DOI: 10.1080/07391102.2020.1815581
Ribonucleocapsid assembly/packaging signals in the genomes of the coronaviruses SARS-CoV and SARS-CoV-2: detection, comparison and implications for therapeutic targeting
Abstract
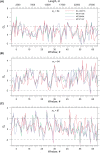
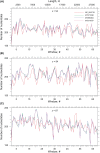
The genomic ssRNA of coronaviruses is packaged within a helical nucleocapsid. Due to transitional symmetry of a helix, weakly specific cooperative interaction between ssRNA and nucleocapsid proteins leads to the natural selection of specific quasi-periodic assembly/packaging signals in the related genomic sequence. Such signals coordinated with the nucleocapsid helical structure were detected and reconstructed in the genomes of the coronaviruses SARS-CoV and SARS-CoV-2. The main period of the signals for both viruses was about 54 nt, that implies 6.75 nt per N protein. The complete coverage of the ssRNA genome of length about 30,000 nt by the nucleocapsid would need 4.4 × 103 N proteins, that makes them the most abundant among the structural proteins. The repertoires of motifs for SARS-CoV and SARS-CoV-2 were divergent but nearly coincided for different isolates of SARS-CoV-2. We obtained the distributions of assembly/packaging signals over the genomes with nonoverlapping windows of width 432 nt. Finally, using the spectral entropy, we compared the load from point mutations and indels during virus age for SARS-CoV and SARS-CoV-2. We found the higher mutational load on SARS-CoV. In this sense, SARS-CoV-2 can be treated as a 'newborn' virus. These observations may be helpful in practical medical applications and are of basic interest. Communicated by Ramaswamy H. Sarma.
Keywords: COVID-19; SARS-CoV; SARS-CoV-2; genome packaging; helical nucleocapsid.
Conflict of interest statement
No potential conflict of interest was reported by the authors.
Figures
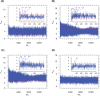
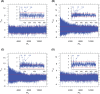
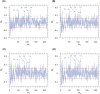
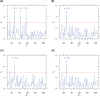
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous
