Camel regulates development of the brain ventricular system
- PMID: 32902807
- PMCID: PMC7904751
- DOI: 10.1007/s00441-020-03270-1
Camel regulates development of the brain ventricular system
Abstract
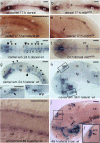
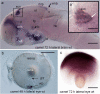
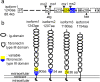
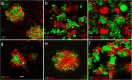
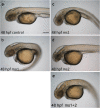
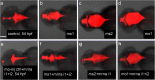
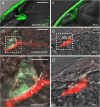
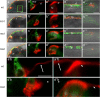
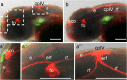
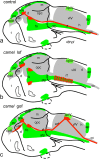
Development of the brain ventricular system of vertebrates and the molecular mechanisms involved are not fully understood. The developmental genes expressed in the elements of the brain ventricular system such as the ependyma and circumventricular organs act as molecular determinants of cell adhesion critical for the formation of brain ventricular system. They control brain development and function, including the flow of cerebrospinal fluid. Here, we describe the novel distantly related member of the zebrafish L1-CAM family of genes-camel. Whereas its maternal transcripts distributed uniformly, the zygotic transcripts demonstrate clearly defined expression patterns, in particular in the axial structures: floor plate, hypochord, and roof plate. camel expresses in several other cell lineages with access to the brain ventricular system, including the midbrain roof plate, subcommissural organ, organum vasculosum lamina terminalis, median eminence, paraventricular organ, flexural organ, and inter-rhombomeric boundaries. This expression pattern suggests a role of Camel in neural development. Several isoforms of Camel generated by differential splicing of exons encoding the sixth fibronectin type III domain enhance cell adhesion differentially. The antisense oligomer morpholino-mediated loss-of-function of Camel affects cell adhesion and causes hydrocephalus and scoliosis manifested via the tail curled down phenotype. The subcommissural organ's derivative-the Reissner fiber-participates in the flow of cerebrospinal fluid. The Reissner fiber fails to form upon morpholino-mediated Camel loss-of-function. The Camel mRNA-mediated gain-of-function causes the Reissner fiber misdirection. This study revealed a link between Chl1a/Camel and Reissner fiber formation, and this supports the idea that CHL1 is one of the scoliosis factors.
Keywords: Ependyma; Flexural organ; Floor plate; Hypochord; Roof plate; Subcommissural organ.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures


Similar articles
-
Expression patterns of dscam and sdk gene paralogs in developing zebrafish retina.Mol Vis. 2018 Jul 19;24:443-458. eCollection 2018. Mol Vis. 2018. PMID: 30078982 Free PMC article.
-
Ambra1 knockdown in zebrafish leads to incomplete development due to severe defects in organogenesis.Autophagy. 2013 Apr;9(4):476-95. doi: 10.4161/auto.23278. Epub 2013 Jan 24. Autophagy. 2013. PMID: 23348054 Free PMC article.
-
The Reissner Fiber Is Highly Dynamic In Vivo and Controls Morphogenesis of the Spine.Curr Biol. 2020 Jun 22;30(12):2353-2362.e3. doi: 10.1016/j.cub.2020.04.015. Epub 2020 May 7. Curr Biol. 2020. PMID: 32386529 Free PMC article.
-
Development of the brain ventricular system from a comparative perspective.Clin Anat. 2023 Mar;36(2):320-334. doi: 10.1002/ca.23994. Epub 2022 Dec 28. Clin Anat. 2023. PMID: 36529666 Review.
-
Involvement of claudins in zebrafish brain ventricle morphogenesis.Ann N Y Acad Sci. 2012 Jun;1257:193-8. doi: 10.1111/j.1749-6632.2012.06507.x. Ann N Y Acad Sci. 2012. PMID: 22671606 Review.
Cited by
-
Brain Barriers and brain fluids research in 2020 and the fluids and barriers of the CNS thematic series on advances in in vitro modeling of the blood-brain barrier and neurovascular unit.Fluids Barriers CNS. 2021 May 21;18(1):24. doi: 10.1186/s12987-021-00258-z. Fluids Barriers CNS. 2021. PMID: 34020685 Free PMC article.
-
Genetic and molecular mechanisms of hydrocephalus.Front Mol Neurosci. 2025 Jan 7;17:1512455. doi: 10.3389/fnmol.2024.1512455. eCollection 2024. Front Mol Neurosci. 2025. PMID: 39839745 Free PMC article. Review.
-
Molecular hallmarks of hydrocephalus.Sci Transl Med. 2025 Jun 4;17(801):eadq1810. doi: 10.1126/scitranslmed.adq1810. Epub 2025 Jun 4. Sci Transl Med. 2025. PMID: 40465691 Free PMC article. Review.
-
Advances in brain barriers and brain fluids research in 2021: great progress in a time of adversity.Fluids Barriers CNS. 2022 Jun 9;19(1):48. doi: 10.1186/s12987-022-00343-x. Fluids Barriers CNS. 2022. PMID: 35681151 Free PMC article.
-
Congenital hydrocephalus: a review of recent advances in genetic etiology and molecular mechanisms.Mil Med Res. 2024 Aug 12;11(1):54. doi: 10.1186/s40779-024-00560-5. Mil Med Res. 2024. PMID: 39135208 Free PMC article. Review.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

