SLC25A51 is a mammalian mitochondrial NAD+ transporter
- PMID: 32906142
- PMCID: PMC7718333
- DOI: 10.1038/s41586-020-2741-7
SLC25A51 is a mammalian mitochondrial NAD+ transporter
Abstract
Mitochondria require nicotinamide adenine dinucleotide (NAD+) to carry out the fundamental processes that fuel respiration and mediate cellular energy transduction. Mitochondrial NAD+ transporters have been identified in yeast and plants1,2, but their existence in mammals remains controversial3-5. Here we demonstrate that mammalian mitochondria can take up intact NAD+, and identify SLC25A51 (also known as MCART1)-an essential6,7 mitochondrial protein of previously unknown function-as a mammalian mitochondrial NAD+ transporter. Loss of SLC25A51 decreases mitochondrial-but not whole-cell-NAD+ content, impairs mitochondrial respiration, and blocks the uptake of NAD+ into isolated mitochondria. Conversely, overexpression of SLC25A51 or SLC25A52 (a nearly identical paralogue of SLC25A51) increases mitochondrial NAD+ levels and restores NAD+ uptake into yeast mitochondria lacking endogenous NAD+ transporters. Together, these findings identify SLC25A51 as a mammalian transporter capable of importing NAD+ into mitochondria.
Figures

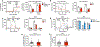







Comment in
-
Maestro of the SereNADe: SLC25A51 Orchestrates Mitochondrial NAD.Trends Biochem Sci. 2021 May;46(5):348-350. doi: 10.1016/j.tibs.2021.02.001. Epub 2021 Feb 19. Trends Biochem Sci. 2021. PMID: 33618948 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

