Multiplex Immunofluorescence in Formalin-Fixed Paraffin-Embedded Tumor Tissue to Identify Single-Cell-Level PI3K Pathway Activation
- PMID: 32913135
- PMCID: PMC7669556
- DOI: 10.1158/1078-0432.CCR-20-2000
Multiplex Immunofluorescence in Formalin-Fixed Paraffin-Embedded Tumor Tissue to Identify Single-Cell-Level PI3K Pathway Activation
Abstract
Purpose: Identifying cancers with high PI3K pathway activity is critical for treatment selection and eligibility into clinical trials of PI3K inhibitors. Assessments of tumor signaling pathway activity need to consider intratumoral heterogeneity and multiple regulatory nodes.
Experimental design: We established a novel, mechanistically informed approach to assessing tumor signaling pathways by quantifying single-cell-level multiplex immunofluorescence using custom algorithms. In a proof-of-concept study, we stained archival formalin-fixed, paraffin-embedded (FFPE) tissue from patients with primary prostate cancer in two prospective cohort studies, the Health Professionals Follow-up Study and the Physicians' Health Study. PTEN, stathmin, and phospho-S6 were quantified on 14 tissue microarrays as indicators of PI3K activation to derive cell-level PI3K scores.
Results: In 1,001 men, 988,254 tumor cells were assessed (median, 743 per tumor; interquartile range, 290-1,377). PI3K scores were higher in tumors with PTEN loss scored by a pathologist, higher Gleason grade, and a new, validated bulk PI3K transcriptional signature. Unsupervised machine-learning approaches resulted in similar clustering. Within-tumor heterogeneity in cell-level PI3K scores was high. During long-term follow-up (median, 15.3 years), rates of progression to metastases and death from prostate cancer were twice as high in the highest quartile of PI3K activation compared with the lowest quartile (hazard ratio, 2.04; 95% confidence interval, 1.13-3.68).
Conclusions: Our novel pathway-focused approach to quantifying single-cell-level immunofluorescence in FFPE tissue identifies prostate tumors with PI3K pathway activation that are more aggressive and may respond to pathway inhibitors.
©2020 American Association for Cancer Research.
Conflict of interest statement
Conflicts of Interest:
P.W. Kantoff is not aware of conflicts of interest. It is his policy to disclose all relationships, which include ownership interest in Context Therapeutics LLC, DRGT, Placon, Seer Biosciences, and Tarveda Therapeutics; he is a company board member for Context Therapeutics LLC, a consultant/advisory board member for BIND Biosciences, Inc., BN Immunotherapeutics, DRGT, GE Healthcare, Janssen, Metamark, New England Research Institutes, Inc., OncoCellMDX, Progenity, Sanofi, Seer Biosciences, Tarveda Therapeutics, and Thermo Fisher, and serves on data safety monitoring boards for Genentech/Roche and Merck. No potential conflicts of interest were disclosed by other authors.
Figures

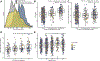
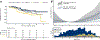

Similar articles
-
Loss of PTEN Expression Detected by Fluorescence Immunohistochemistry Predicts Lethal Prostate Cancer in Men Treated with Prostatectomy.Eur Urol Oncol. 2019 Sep;2(5):475-482. doi: 10.1016/j.euo.2018.09.003. Epub 2018 Oct 4. Eur Urol Oncol. 2019. PMID: 31411988
-
Poor prognosis in carcinoma is associated with a gene expression signature of aberrant PTEN tumor suppressor pathway activity.Proc Natl Acad Sci U S A. 2007 May 1;104(18):7564-9. doi: 10.1073/pnas.0702507104. Epub 2007 Apr 23. Proc Natl Acad Sci U S A. 2007. PMID: 17452630 Free PMC article.
-
Coffee, Phosphoinositide 3-Kinase Signaling Pathway, and Prostate Cancer: A Prospective Study in the Health Professionals Follow-Up Study.J Acad Nutr Diet. 2025 Jan;125(1):90-98.e5. doi: 10.1016/j.jand.2024.07.001. Epub 2024 Jul 4. J Acad Nutr Diet. 2025. PMID: 38971221
-
Deregulated PTEN/PI3K/AKT/mTOR signaling in prostate cancer: Still a potential druggable target?Biochim Biophys Acta Mol Cell Res. 2020 Sep;1867(9):118731. doi: 10.1016/j.bbamcr.2020.118731. Epub 2020 Apr 29. Biochim Biophys Acta Mol Cell Res. 2020. PMID: 32360668 Review.
-
Targeting the PI3K/Akt pathway in prostate cancer: challenges and opportunities (review).Int J Oncol. 2014 Nov;45(5):1793-801. doi: 10.3892/ijo.2014.2601. Epub 2014 Aug 14. Int J Oncol. 2014. PMID: 25120209 Review.
Cited by
-
MTFR2-Mediated Fission Drives Fatty Acid and Mitochondrial Co-Transfer from Hepatic Stellate Cells to Tumor Cells Fueling Oncogenesis.Adv Sci (Weinh). 2025 Jun;12(23):e2416419. doi: 10.1002/advs.202416419. Epub 2025 May 14. Adv Sci (Weinh). 2025. PMID: 40365837 Free PMC article.
-
A Personalized Genomics Approach of the Prostate Cancer.Cells. 2021 Jun 30;10(7):1644. doi: 10.3390/cells10071644. Cells. 2021. PMID: 34209090 Free PMC article.
-
NaroNet: Discovery of tumor microenvironment elements from highly multiplexed images.Med Image Anal. 2022 May;78:102384. doi: 10.1016/j.media.2022.102384. Epub 2022 Feb 14. Med Image Anal. 2022. PMID: 35217454 Free PMC article.
-
Extent, impact, and mitigation of batch effects in tumor biomarker studies using tissue microarrays.Elife. 2021 Dec 23;10:e71265. doi: 10.7554/eLife.71265. Elife. 2021. PMID: 34939926 Free PMC article.
-
Distinct mesenchymal cell states mediate prostate cancer progression.Nat Commun. 2024 Jan 8;15(1):363. doi: 10.1038/s41467-023-44210-1. Nat Commun. 2024. PMID: 38191471 Free PMC article.
References
-
- Armstrong AJ, Halabi S, Healy P, Alumkal JJ, Winters C, Kephart J, et al. Phase II trial of the PI3 kinase inhibitor buparlisib (BKM-120) with or without enzalutamide in men with metastatic castration resistant prostate cancer. Eur J Cancer 2017;81:228–36 doi 10.1016/j.ejca.2017.02.030. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

