The Heterotrophic Bacterium Cupriavidus pinatubonensis JMP134 Oxidizes Sulfide to Sulfate with Thiosulfate as a Key Intermediate
- PMID: 32917752
- PMCID: PMC7642071
- DOI: 10.1128/AEM.01835-20
The Heterotrophic Bacterium Cupriavidus pinatubonensis JMP134 Oxidizes Sulfide to Sulfate with Thiosulfate as a Key Intermediate
Abstract
Heterotrophic bacteria actively participate in the biogeochemical cycle of sulfur on Earth. The heterotrophic bacterium Cupriavidus pinatubonensis JMP134 contains several enzymes involved in sulfur oxidation, but how these enzymes work together to oxidize sulfide in the bacterium has not been studied. Using gene-deletion and whole-cell assays, we determined that the bacterium uses sulfide:quinone oxidoreductase to oxidize sulfide to polysulfide, which is further oxidized to sulfite by persulfide dioxygenase. Sulfite spontaneously reacts with polysulfide to produce thiosulfate. The sulfur-oxidizing (Sox) system oxidizes thiosulfate to sulfate. Flavocytochrome c sulfide dehydrogenase enhances thiosulfate oxidation by the Sox system but couples with the Sox system for sulfide oxidation to sulfate in the absence of sulfide:quinone oxidoreductase. Thus, C. pinatubonensis JMP134 contains a main pathway and a contingent pathway for sulfide oxidation.IMPORTANCE We establish a new pathway of sulfide oxidation with thiosulfate as a key intermediate in Cupriavidus pinatubonensis JMP134. The bacterium mainly oxidizes sulfide by using sulfide:quinone oxidoreductase, persulfide dioxygenase, and the Sox system with thiosulfate as a key intermediate. Although the purified and reconstituted Sox system oxidizes sulfide, its rate of sulfide oxidation in C. pinatubonensis JMP134 is too low to be physiologically relevant. The findings reveal how these sulfur-oxidizing enzymes participate in sulfide oxidation in a single bacterium.
Keywords: heterotrophic bacteria; sulfane sulfur; sulfate; sulfate reduction; sulfide; sulfur oxidation pathway; thiosulfate.
Copyright © 2020 American Society for Microbiology.
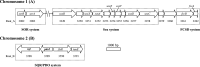
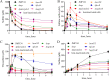
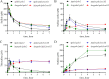
Figures
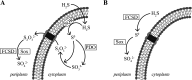
Similar articles
-
Cytoplasmic Localization of Sulfide:Quinone Oxidoreductase and Persulfide Dioxygenase of Cupriavidus pinatubonensis JMP134.Appl Environ Microbiol. 2017 Nov 16;83(23):e01820-17. doi: 10.1128/AEM.01820-17. Print 2017 Dec 1. Appl Environ Microbiol. 2017. PMID: 28939597 Free PMC article.
-
Cupriavidus pinatubonensis JMP134 Alleviates Sulfane Sulfur Toxicity after the Loss of Sulfane Dehydrogenase through Oxidation by Persulfide Dioxygenase and Hydrogen Sulfide Release.Metabolites. 2023 Feb 2;13(2):218. doi: 10.3390/metabo13020218. Metabolites. 2023. PMID: 36837837 Free PMC article.
-
Cupriavidus necator H16 Uses Flavocytochrome c Sulfide Dehydrogenase To Oxidize Self-Produced and Added Sulfide.Appl Environ Microbiol. 2017 Oct 31;83(22):e01610-17. doi: 10.1128/AEM.01610-17. Print 2017 Nov 15. Appl Environ Microbiol. 2017. PMID: 28864655 Free PMC article.
-
[Oxidation of inorganic sulfur compounds by obligatory organotrophic bacteria].Mikrobiologiia. 2003 Nov-Dec;72(6):725-39. Mikrobiologiia. 2003. PMID: 14768537 Review. Russian.
-
Inorganic sulfur oxidizing system in green sulfur bacteria.Photosynth Res. 2010 Jun;104(2-3):163-76. doi: 10.1007/s11120-010-9531-2. Epub 2010 Feb 9. Photosynth Res. 2010. PMID: 20143161 Review.
Cited by
-
Mechanism of Intracellular Elemental Sulfur Oxidation in Beggiatoa leptomitoformis, Where Persulfide Dioxygenase Plays a Key Role.Int J Mol Sci. 2024 Oct 11;25(20):10962. doi: 10.3390/ijms252010962. Int J Mol Sci. 2024. PMID: 39456744 Free PMC article.
-
The Mechanisms of Thiosulfate Toxicity against Saccharomyces cerevisiae.Antioxidants (Basel). 2021 Apr 22;10(5):646. doi: 10.3390/antiox10050646. Antioxidants (Basel). 2021. PMID: 33922196 Free PMC article.
-
The critical roles of propanethiol oxidoreductase and sulfide-quinone oxidoreductase in the propanethiol catabolism pathway in Pseudomonas putida S-1.Appl Environ Microbiol. 2024 Feb 21;90(2):e0195923. doi: 10.1128/aem.01959-23. Epub 2024 Jan 9. Appl Environ Microbiol. 2024. PMID: 38193681 Free PMC article.
-
OxyR-regulated T6SS functions in coordination with siderophore to resist oxidative stress.Microbiol Spectr. 2024 Feb 6;12(2):e0323123. doi: 10.1128/spectrum.03231-23. Epub 2024 Jan 8. Microbiol Spectr. 2024. PMID: 38189330 Free PMC article.
-
Sulfane Sulfur Posttranslationally Modifies the Global Regulator AdpA to Influence Actinorhodin Production and Morphological Differentiation of Streptomyces coelicolor.mBio. 2022 Jun 28;13(3):e0386221. doi: 10.1128/mbio.03862-21. Epub 2022 Apr 25. mBio. 2022. PMID: 35467418 Free PMC article.
References
Publication types
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

