Present and Emerging Methodologies in Cryo-EM Single-Particle Analysis
- PMID: 32919493
- PMCID: PMC7567993
- DOI: 10.1016/j.bpj.2020.08.027
Present and Emerging Methodologies in Cryo-EM Single-Particle Analysis
Abstract
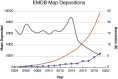
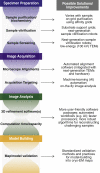
Over the past decade, technical and methodological improvements in cryogenic electron microscopy (cryo-EM) single-particle analysis have enabled routine high-resolution structural analyses of biological macromolecules, resulting in a flood of new molecular insights into protracted biological questions. However, despite the tremendous progress and success of the field in recent years, opportunities for improvement remain in various aspects of the cryo-EM single-particle analysis workflow (e.g., sample preparation, image acquisition and processing, and structure validation). Here, we review recent advances that have contributed to the principal methods in cryo-EM and identify persisting challenges and bottlenecks that will require further methodological and hardware development.
Copyright © 2020 Biophysical Society. Published by Elsevier Inc. All rights reserved.
Figures
Similar articles
-
Sample preparation of biological macromolecular assemblies for the determination of high-resolution structures by cryo-electron microscopy.Microscopy (Oxf). 2016 Feb;65(1):23-34. doi: 10.1093/jmicro/dfv367. Epub 2015 Dec 15. Microscopy (Oxf). 2016. PMID: 26671943 Review.
-
Retrospect and Prospect of Single Particle Cryo-Electron Microscopy: The Class of Integral Membrane Proteins as an Example.J Chem Inf Model. 2020 May 26;60(5):2448-2457. doi: 10.1021/acs.jcim.9b01015. Epub 2020 Mar 20. J Chem Inf Model. 2020. PMID: 32163280 Review.
-
Single-particle cryo-electron microscopy of macromolecular complexes.Microscopy (Oxf). 2016 Feb;65(1):9-22. doi: 10.1093/jmicro/dfv366. Epub 2015 Nov 25. Microscopy (Oxf). 2016. PMID: 26611544 Free PMC article. Review.
-
Miniaturizing EM Sample Preparation: Opportunities, Challenges, and "Visual Proteomics".Proteomics. 2018 Mar;18(5-6):e1700176. doi: 10.1002/pmic.201700176. Proteomics. 2018. PMID: 29441686 Review.
-
Approaches to altering particle distributions in cryo-electron microscopy sample preparation.Acta Crystallogr D Struct Biol. 2018 Jun 1;74(Pt 6):560-571. doi: 10.1107/S2059798318006496. Epub 2018 May 18. Acta Crystallogr D Struct Biol. 2018. PMID: 29872006 Free PMC article. Review.
Cited by
-
Insights into the flexibility of the domain-linking loop in actinobacterial coproheme decarboxylase through structures and molecular dynamics simulations.Protein Sci. 2025 Feb;34(2):e70027. doi: 10.1002/pro.70027. Protein Sci. 2025. PMID: 39865384 Free PMC article.
-
Integrative approaches in cryogenic electron microscopy: Recent advances in structural biology and future perspectives.iScience. 2021 Jan 9;24(2):102044. doi: 10.1016/j.isci.2021.102044. eCollection 2021 Feb 19. iScience. 2021. PMID: 33532719 Free PMC article. Review.
-
3D variability analysis reveals a hidden conformational change controlling ammonia transport in human asparagine synthetase.Nat Commun. 2024 Dec 3;15(1):10538. doi: 10.1038/s41467-024-54912-9. Nat Commun. 2024. PMID: 39627226 Free PMC article.
-
High-Throughput Identification of Crystalline Natural Products from Crude Extracts Enabled by Microarray Technology and microED.ACS Cent Sci. 2023 Dec 20;10(1):176-183. doi: 10.1021/acscentsci.3c01365. eCollection 2024 Jan 24. ACS Cent Sci. 2023. PMID: 38292598 Free PMC article.
-
3D Variability Analysis Reveals a Hidden Conformational Change Controlling Ammonia Transport in Human Asparagine Synthetase.bioRxiv [Preprint]. 2024 Sep 5:2023.05.16.541009. doi: 10.1101/2023.05.16.541009. bioRxiv. 2024. Update in: Nat Commun. 2024 Dec 3;15(1):10538. doi: 10.1038/s41467-024-54912-9. PMID: 37292727 Free PMC article. Updated. Preprint.
References
-
- Yan C., Wan R., Shi Y. Structure of a yeast activated spliceosome at 3.5 Å resolution. Science. 2016;353:904–911. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

