Availability of Nitrite and Nitrate as Electron Acceptors Modulates Anaerobic Toluene-Degrading Communities in Aquifer Sediments
- PMID: 32922372
- PMCID: PMC7456981
- DOI: 10.3389/fmicb.2020.01867
Availability of Nitrite and Nitrate as Electron Acceptors Modulates Anaerobic Toluene-Degrading Communities in Aquifer Sediments
Abstract
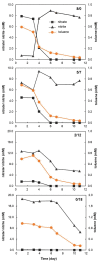
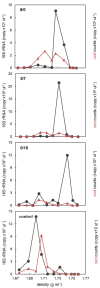
Microorganisms are essential in the degradation of environmental pollutants. Aromatic hydrocarbons, e.g., benzene, toluene, ethylbenzene, and xylene (BTEX), are common aquifer contaminants, whose degradation in situ is often limited by the availability of electron acceptors. It is clear that different electron acceptors such as nitrate, iron, or sulfate support the activity of distinct degraders. However, this has not been demonstrated for the availability of nitrate vs. nitrite, both of which can be respired in reductive nitrogen cycling. Here via DNA-stable isotope probing, we report that nitrate and nitrite provided as electron acceptors in different concentrations and ratios not only modulated the microbial communities responsible for toluene degradation but also influenced how nitrate reduction proceeded. Zoogloeaceae members, mainly Azoarcus spp., were the key toluene degraders with nitrate-only, or both nitrate and nitrite as electron acceptors. In addition, a shift within Azoarcus degrader populations was observed on the amplicon sequence variant (ASV) level depending on electron acceptor ratios. In contrast, members of the Sphingomonadales were likely the most active toluene degraders when only nitrite was provided. Nitrate reduction did not proceed beyond nitrite in the nitrate-only treatment, while it continued when nitrite was initially also present in the microcosms. Likely, this was attributed to the fact that different microbial communities were stimulated and active in different microcosms. Together, these findings demonstrate that the availability of nitrate and nitrite can define degrader community selection and N-reduction outcomes. It also implies that nitrate usage efficiency in bioremediation could possibly be enhanced by an initial co-supply of nitrite, via modulating the active degrader communities.
Keywords: Azoarcus; bioremediation; oxygenic denitrification; stable isotope probing; toluene degradation.
Copyright © 2020 Zhu, Friedrich, Wang, Táncsics and Lueders.
Figures

Similar articles
-
A strategy for aromatic hydrocarbon bioremediation under anaerobic conditions and the impacts of ethanol: a microcosm study.J Contam Hydrol. 2008 Feb 19;96(1-4):17-31. doi: 10.1016/j.jconhyd.2007.09.006. Epub 2007 Sep 29. J Contam Hydrol. 2008. PMID: 17964687
-
Electron acceptors determine the BTEX degradation capacity of anaerobic microbiota via regulating the microbial community.Environ Res. 2022 Dec;215(Pt 3):114420. doi: 10.1016/j.envres.2022.114420. Epub 2022 Sep 24. Environ Res. 2022. PMID: 36167116
-
Microbial community of a gasworks aquifer and identification of nitrate-reducing Azoarcus and Georgfuchsia as key players in BTEX degradation.Water Res. 2018 Apr 1;132:146-157. doi: 10.1016/j.watres.2017.12.040. Epub 2017 Dec 20. Water Res. 2018. PMID: 29324294
-
The use of nucleic acid based stable isotope probing to identify the microorganisms responsible for anaerobic benzene and toluene biodegradation.J Microbiol Methods. 2011 May;85(2):83-91. doi: 10.1016/j.mimet.2011.02.011. Epub 2011 Feb 26. J Microbiol Methods. 2011. PMID: 21356251 Review.
-
Metabolism of alkylbenzenes, alkanes, and other hydrocarbons in anaerobic bacteria.Biodegradation. 2000;11(2-3):85-105. doi: 10.1023/a:1011122631799. Biodegradation. 2000. PMID: 11440245 Review.
Cited by
-
Alterations in bacterial structure and function in seawater due to Mytilus coruscus farming: implications for sustainable aquaculture management.Front Microbiol. 2025 Apr 3;16:1567340. doi: 10.3389/fmicb.2025.1567340. eCollection 2025. Front Microbiol. 2025. PMID: 40248425 Free PMC article.
-
Microaerobic biodegradation of aromatic hydrocarbon mixtures: strategies for efficient nitrate and oxygen dosage.Appl Microbiol Biotechnol. 2025 Jan 16;109(1):9. doi: 10.1007/s00253-024-13388-9. Appl Microbiol Biotechnol. 2025. PMID: 39821078 Free PMC article.
-
Microbial production of toluene in oxygen minimum zone waters in the Humboldt Current System off Chile.Sci Rep. 2022 Jun 23;12(1):10669. doi: 10.1038/s41598-022-14103-2. Sci Rep. 2022. PMID: 35739129 Free PMC article.
-
Unravelling the genetic and functional diversity of dominant bacterial communities involved in manure co-composting bioremediation of complex crude oil waste sludge.Heliyon. 2022 Feb 11;8(2):e08945. doi: 10.1016/j.heliyon.2022.e08945. eCollection 2022 Feb. Heliyon. 2022. PMID: 35243067 Free PMC article.
-
Reconnaissance of Oxygenic Denitrifiers in Agriculturally Impacted Soils.mSphere. 2023 Jun 22;8(3):e0057122. doi: 10.1128/msphere.00571-22. Epub 2023 Apr 5. mSphere. 2023. PMID: 37017537 Free PMC article.
References
-
- Boden R., Hutt L. P., Rae A. W. (2017). Reclassification of Thiobacillus aquaesulis (Wood & Kelly, 1995) as Annwoodia aquaesulis gen. nov., comb. nov., transfer of Thiobacillus (Beijerinck, 1904) from the Hydrogenophilales to the Nitrosomonadales, proposal of Hydrogenophilalia class. nov. within the ‘Proteobacteria’, and four new families within the orders Nitrosomonadales and Rhodocyclales. Int. J. Syst. Evol. Microbiol. 67, 1191–1205. 10.1099/ijsem.0.001927, PMID: - DOI - PubMed
-
- Bradford L. M., Vestergaard G., Táncsics A., Zhu B., Schloter M., Lueders T. (2018). Transcriptome-stable isotope probing provides targeted functional and taxonomic insights into microaerobic pollutant-degrading aquifer microbiota. Front. Microbiol. 9:2696. 10.3389/fmicb.2018.02696, PMID: - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

