Multidrug Resistant Acinetobacter Isolates Release Resistance Determinants Through Contact-Dependent Killing and Bacteriophage Lysis
- PMID: 32922376
- PMCID: PMC7456956
- DOI: 10.3389/fmicb.2020.01918
Multidrug Resistant Acinetobacter Isolates Release Resistance Determinants Through Contact-Dependent Killing and Bacteriophage Lysis
Abstract
Antimicrobial resistance is an ancient bacterial defense mechanism that has rapidly spread due to the frequent use of antibiotics for disease treatment and livestock growth promotion. We are becoming increasingly aware that pathogens, such as members of the genus Acinetobacter, are precipitously evolving drug resistances through multiple mechanisms, including the acquisition of antibiotic resistance genes. In this study, we isolated three multidrug resistant Acinetobacter species from birds on a free-range farm. Acinetobacter radioresistens, Acinetobacter lwoffii, and Acinetobacter johnsonii were isolated from hens, turkeys and ducks and were resistant to 14 clinically relevant antibiotics, including several listed by the World Health Organization as essential medicines. Co-culturing any of the three Acinetobacter species with Acinetobacter baumannii resulted in contact-dependent release of intact resistance determinants. We also isolated several lytic bacteriophages and selected two of these phages to be included in this study based on differences in plaquing characteristics, nucleic acid content and viral morphology. Both phages released host DNA, including antibiotic resistance genes during cell lysis and we demonstrated that these resistance determinants were transferable to a naïve strain of Escherichia coli. This study demonstrates that contact-dependent competition between bacterial species can readily contribute to DNA release into the environment, including antibiotic resistance determinants. We also highlight that the constant lysis and turnover of bacterial populations during the natural lifecycle of a lytic bacteriophage is an underappreciated mechanism for the liberation of DNA and subsequent genetic exchange.
Keywords: Acinetobacter; bacteriophages; contact-dependent killing; environmental isolates; gene transfer; multidrug resistance.
Copyright © 2020 Crippen, Jr., Sanchez and Szymanski.
Figures
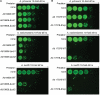


References
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

