Structure of the bacterial ribosome at 2 Å resolution
- PMID: 32924932
- PMCID: PMC7550191
- DOI: 10.7554/eLife.60482
Structure of the bacterial ribosome at 2 Å resolution
Abstract
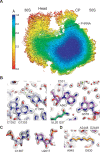
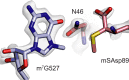
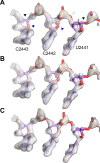
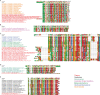
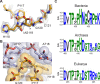
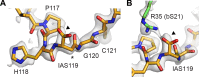
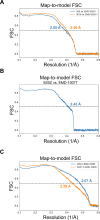
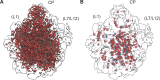
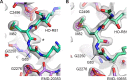
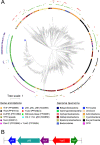
Using cryo-electron microscopy (cryo-EM), we determined the structure of the Escherichia coli 70S ribosome with a global resolution of 2.0 Å. The maps reveal unambiguous positioning of protein and RNA residues, their detailed chemical interactions, and chemical modifications. Notable features include the first examples of isopeptide and thioamide backbone substitutions in ribosomal proteins, the former likely conserved in all domains of life. The maps also reveal extensive solvation of the small (30S) ribosomal subunit, and interactions with A-site and P-site tRNAs, mRNA, and the antibiotic paromomycin. The maps and models of the bacterial ribosome presented here now allow a deeper phylogenetic analysis of ribosomal components including structural conservation to the level of solvation. The high quality of the maps should enable future structural analyses of the chemical basis for translation and aid the development of robust tools for cryo-EM structure modeling and refinement.
Keywords: Cryo-EM; E. coli; aminoglycoside; antibiotics; molecular biophysics; post-transcriptional modifications; post-translational modifications; ribosome; structural biology.
Plain language summary
Inside cells, proteins are produced by complex molecular machines called ribosomes. Techniques that allow scientists to visualize ribosomes at the atomic level, such as cryogenic electron microscopy (cryo-EM), help shed light on the structure of these molecular machines, revealing details of how they build proteins. Understanding how ribosomes work has many benefits, including the development of new antibiotics that can kill bacteria without affecting animal cells. Watson et al. used cryo-EM techniques with increased resolution to examine the ribosomes of the bacterium Escherichia coli in a higher level of detail than has been seen before. The results revealed two chemical modifications in proteins that form the ribosome that had not been observed in ribosomes previously. Additionally, a protein segment with a previously undescribed structure was identified close to the site where the ribosome reads the genetic instructions needed to make proteins. Further genetic analyses suggested these structures are in many related species, and may play important roles in how the ribosome works. Watson et al. were also able to see how paromomycin, an antibiotic used to treat parasitic infections, is positioned in the ribosome. The antibiotic interacts with a site near where the genetic code is read out, which might explain why certain changes to the antibiotic can interfere with its potency. Finally, the new ribosome structure reveals thousands of water molecules and metal ions that help keep the ribosome together as it produces proteins. This study shows the value of advances in cryo-EM technology and illustrates the importance of applying these techniques to other cell components. The results also reveal details of the ribosome useful for further research into this essential molecular machine.
© 2020, Watson et al.
Conflict of interest statement
ZW, FW, RM, OA, AS, JB, JC No competing interests declared
Figures







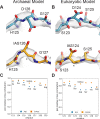
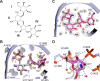
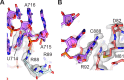
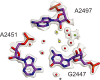

Similar articles
-
Structure of the E. coli ribosome-EF-Tu complex at <3 Å resolution by Cs-corrected cryo-EM.Nature. 2015 Apr 23;520(7548):567-70. doi: 10.1038/nature14275. Epub 2015 Feb 23. Nature. 2015. PMID: 25707802
-
Direct three-dimensional localization and positive identification of RNA helices within the ribosome by means of genetic tagging and cryo-electron microscopy.Structure. 1999 Dec 15;7(12):1567-73. doi: 10.1016/s0969-2126(00)88347-1. Structure. 1999. PMID: 10647187
-
A method for differentiating proteins from nucleic acids in intermediate-resolution density maps: cryo-electron microscopy defines the quaternary structure of the Escherichia coli 70S ribosome.Structure. 2000 Sep 15;8(9):937-48. doi: 10.1016/s0969-2126(00)00185-4. Structure. 2000. PMID: 10986461
-
Cryo-electron microscopy as an investigative tool: the ribosome as an example.Bioessays. 2001 Aug;23(8):725-32. doi: 10.1002/bies.1102. Bioessays. 2001. PMID: 11494321 Review.
-
Ribosome dynamics: insights from atomic structure modeling into cryo-electron microscopy maps.Annu Rev Biophys Biomol Struct. 2006;35:299-317. doi: 10.1146/annurev.biophys.35.040405.101950. Annu Rev Biophys Biomol Struct. 2006. PMID: 16689638 Review.
Cited by
-
Analysis and comparison of electron radiation damage assessments in Cryo-EM by single particle analysis and micro-crystal electron diffraction.Front Mol Biosci. 2022 Oct 5;9:988928. doi: 10.3389/fmolb.2022.988928. eCollection 2022. Front Mol Biosci. 2022. PMID: 36275612 Free PMC article. Review.
-
Mechanism of Inhibition of the Reproduction of SARS-CoV-2 and Ebola Viruses by Remdesivir.Biochemistry. 2021 Jun 22;60(24):1869-1875. doi: 10.1021/acs.biochem.1c00292. Epub 2021 Jun 10. Biochemistry. 2021. PMID: 34110129 Free PMC article.
-
Minimization of the E. coli ribosome, aided and optimized by community science.Nucleic Acids Res. 2024 Feb 9;52(3):1027-1042. doi: 10.1093/nar/gkad1254. Nucleic Acids Res. 2024. PMID: 38214230 Free PMC article.
-
Cryo-EM captures early ribosome assembly in action.Nat Commun. 2023 Feb 17;14(1):898. doi: 10.1038/s41467-023-36607-9. Nat Commun. 2023. PMID: 36797249 Free PMC article.
-
Incorporating thioamides into proteins by native chemical ligation.Methods Enzymol. 2021;656:295-339. doi: 10.1016/bs.mie.2021.04.011. Epub 2021 May 7. Methods Enzymol. 2021. PMID: 34325791 Free PMC article.
References
-
- Al-Shayeb B, Sachdeva R, Chen LX, Ward F, Munk P, Devoto A, Castelle CJ, Olm MR, Bouma-Gregson K, Amano Y, He C, Méheust R, Brooks B, Thomas A, Lavy A, Matheus-Carnevali P, Sun C, Goltsman DSA, Borton MA, Sharrar A, Jaffe AL, Nelson TC, Kantor R, Keren R, Lane KR, Farag IF, Lei S, Finstad K, Amundson R, Anantharaman K, Zhou J, Probst AJ, Power ME, Tringe SG, Li WJ, Wrighton K, Harrison S, Morowitz M, Relman DA, Doudna JA, Lehours AC, Warren L, Cate JHD, Santini JM, Banfield JF. Clades of huge phages from across earth's ecosystems. Nature. 2020;578:425–431. doi: 10.1038/s41586-020-2007-4. - DOI - PMC - PubMed
-
- Anantharaman K, Brown CT, Hug LA, Sharon I, Castelle CJ, Probst AJ, Thomas BC, Singh A, Wilkins MJ, Karaoz U, Brodie EL, Williams KH, Hubbard SS, Banfield JF. Thousands of microbial genomes shed light on interconnected biogeochemical processes in an aquifer system. Nature Communications. 2016;7:13219. doi: 10.1038/ncomms13219. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

