Type 2-high asthma is associated with a specific indoor mycobiome and microbiome
- PMID: 32926879
- PMCID: PMC7486598
- DOI: 10.1016/j.jaci.2020.08.035
Type 2-high asthma is associated with a specific indoor mycobiome and microbiome
Abstract
Background: The links between microbial environmental exposures and asthma are well documented, but no study has combined deep sequencing results from pulmonary and indoor microbiomes of patients with asthma with spirometry, clinical, and endotype parameters.
Objective: The goal of this study was to investigate the links between indoor microbial exposures and pulmonary microbial communities and to document the role of microbial exposures on inflammatory and clinical outcomes of patients with severe asthma (SA).
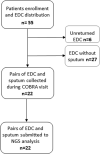
Methods: A total of 55 patients with SA from the national Cohort of Bronchial Obstruction and Asthma cohort were enrolled for analyzing their indoor microbial flora through the use of electrostatic dust collectors (EDCs). Among these patients, 22 were able to produce sputum during "stable" or pulmonary "exacerbation" periods and had complete pairs of EDC and sputum samples, both collected and analyzed. We used amplicon targeted metagenomics to compare microbial communities from EDC and sputum samples of patients according to type 2 (T2)-asthma endotypes.
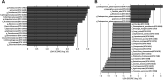
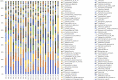
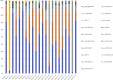
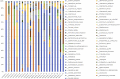
Results: Compared with patients with T2-low SA, patients with T2-high SA exhibited an increase in bacterial α-diversity and a decrease in fungal α-diversity of their indoor microbial florae, the latter being significantly correlated with fraction of exhaled nitric oxide levels. The β-diversity of the EDC mycobiome clustered significantly according to T2 endotypes. Moreover, the proportion of fungal taxa in common between the sputum and EDC samples was significantly higher when patients exhibited acute exacerbation.
Conclusion: These results illustrated, for the first time, a potential association between the indoor mycobiome and clinical features of patients with SA, which should renew interest in deciphering the interactions between indoor environment, fungi, and host in asthma.
Keywords: Feno; Indoor environment; microbiome; mycobiome; severe asthma; type 2 asthma.
Copyright © 2020 American Academy of Allergy, Asthma & Immunology. Published by Elsevier Inc. All rights reserved.
Figures




Comment in
-
Compositional similarity between indoor and human sputum microbiome.J Allergy Clin Immunol. 2021 Feb;147(2):779. doi: 10.1016/j.jaci.2020.10.026. Epub 2020 Dec 1. J Allergy Clin Immunol. 2021. PMID: 33276992 No abstract available.
-
Reply.J Allergy Clin Immunol. 2021 Feb;147(2):779-780. doi: 10.1016/j.jaci.2020.10.021. Epub 2020 Dec 1. J Allergy Clin Immunol. 2021. PMID: 33276993 No abstract available.
Similar articles
-
Characterization of the bacterial and fungal microbiome in indoor dust and outdoor air samples: a pilot study.Environ Sci Process Impacts. 2016 Jun 15;18(6):713-24. doi: 10.1039/c5em00639b. Epub 2016 May 23. Environ Sci Process Impacts. 2016. PMID: 27213188 Free PMC article.
-
The mycobiomes and bacteriomes of sputum, saliva, and home dust.Indoor Air. 2021 Mar;31(2):357-368. doi: 10.1111/ina.12750. Epub 2020 Oct 7. Indoor Air. 2021. PMID: 32969526
-
Fungal and Bacterial Communities in Indoor Dust Follow Different Environmental Determinants.PLoS One. 2016 Apr 21;11(4):e0154131. doi: 10.1371/journal.pone.0154131. eCollection 2016. PLoS One. 2016. PMID: 27100967 Free PMC article.
-
Comparison of Air Impaction and Electrostatic Dust Collector Sampling Methods to Assess Airborne Fungal Contamination in Public Buildings.Ann Occup Hyg. 2016 Mar;60(2):161-75. doi: 10.1093/annhyg/mev075. Epub 2015 Oct 21. Ann Occup Hyg. 2016. PMID: 26491105
-
Indoor microbiome and allergic diseases: From theoretical advances to prevention strategies.Eco Environ Health. 2022 Oct 17;1(3):133-146. doi: 10.1016/j.eehl.2022.09.002. eCollection 2022 Sep. Eco Environ Health. 2022. PMID: 38075599 Free PMC article. Review.
Cited by
-
Lung Mycobiota α-Diversity Is Linked to Severity in Critically Ill Patients with Acute Exacerbation of Chronic Obstructive Pulmonary Disease.Microbiol Spectr. 2023 Mar 28;11(2):e0506222. doi: 10.1128/spectrum.05062-22. Online ahead of print. Microbiol Spectr. 2023. PMID: 36976010 Free PMC article.
-
Microbiome Research and Multi-Omics Integration for Personalized Medicine in Asthma.J Pers Med. 2021 Dec 5;11(12):1299. doi: 10.3390/jpm11121299. J Pers Med. 2021. PMID: 34945771 Free PMC article. Review.
-
Gut Microbiota and Mycobiota Evolution Is Linked to Memory Improvement after Bariatric Surgery in Obese Patients: A Pilot Study.Nutrients. 2021 Nov 13;13(11):4061. doi: 10.3390/nu13114061. Nutrients. 2021. PMID: 34836316 Free PMC article.
-
Relative Humidity and Its Impact on the Immune System and Infections.Int J Mol Sci. 2023 May 29;24(11):9456. doi: 10.3390/ijms24119456. Int J Mol Sci. 2023. PMID: 37298409 Free PMC article. Review.
-
New insights into the pathophysiology and therapeutic targets of asthma and comorbid chronic rhinosinusitis with or without nasal polyposis.Clin Sci (Lond). 2023 May 18;137(9):727-753. doi: 10.1042/CS20190281. Clin Sci (Lond). 2023. PMID: 37199256 Free PMC article. Review.
References
-
- Global Initiative for Asthma GINA annual report. http://ginasthma.org/archived-reports/ Accessed June 2019.
-
- Chung K.F., Wenzel S.E., Brozek J.L., Bush A., Castro M., Sterk P.J., et al. International ERS/ATS guidelines on definition, evaluation and treatment of severe asthma. Eur Respir J. 2014;43:343–373. - PubMed
-
- Li N., Qiu R., Yang Z., Li J., Chung K.F., Zhong N., et al. Sputum microbiota in severe asthma patients: relationship to eosinophilic inflammation. Respir Med. 2017;131:192–198. - PubMed

