The sustained expression of Cas9 targeting toxic RNAs reverses disease phenotypes in mouse models of myotonic dystrophy type 1
- PMID: 32929188
- PMCID: PMC8241012
- DOI: 10.1038/s41551-020-00607-7
The sustained expression of Cas9 targeting toxic RNAs reverses disease phenotypes in mouse models of myotonic dystrophy type 1
Abstract
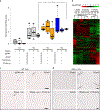
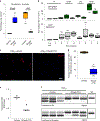
Myotonic dystrophy type I (DM1) is a multisystemic autosomal-dominant inherited human disorder that is caused by CTG microsatellite repeat expansions (MREs) in the 3' untranslated region of DMPK. Toxic RNAs expressed from such repetitive sequences can be eliminated using CRISPR-mediated RNA targeting, yet evidence of its in vivo efficacy and durability is lacking. Here, using adult and neonatal mouse models of DM1, we show that intramuscular or systemic injections of adeno-associated virus (AAV) vectors encoding nuclease-dead Cas9 and a single-guide RNA targeting CUG repeats results in the expression of the RNA-targeting Cas9 for up to three months, redistribution of the RNA-splicing protein muscleblind-like splicing regulator 1, elimination of foci of toxic RNA, reversal of splicing biomarkers and amelioration of myotonia. The sustained reversal of DM1 phenotypes provides further support that RNA-targeting Cas9 is a viable strategy for treating DM1 and other MRE-associated diseases.
Conflict of interest statement
Competing interests
G.W.Y. is a cofounder, member of the board of directors, equity holder and paid consultant of Locanabio. D.A.N. is a cofounder and an equity holder of Locanabio. R.B. is an equity holder and employee of Locanabio. M.S.S. is an equity holder of Locanabio and a Scientific Advisory Board member of Skyhawk Therapeutics. The terms of this arrangement have been reviewed and approved by the University of California San Diego and the University of Florida, Gainesville in accordance with their conflict of interest policies. The other authors declare no other competing interests.
Figures



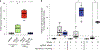
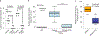
Comment in
-
Cas9 targeting of toxic foci of RNA repeats.Nat Biomed Eng. 2021 Feb;5(2):130-131. doi: 10.1038/s41551-021-00688-y. Nat Biomed Eng. 2021. PMID: 33580228 No abstract available.
Similar articles
-
Muscle-specific gene editing improves molecular and phenotypic defects in a mouse model of myotonic dystrophy type 1.Clin Transl Med. 2025 Feb;15(2):e70227. doi: 10.1002/ctm2.70227. Clin Transl Med. 2025. PMID: 39956955 Free PMC article.
-
Genome Editing of Expanded CTG Repeats within the Human DMPK Gene Reduces Nuclear RNA Foci in the Muscle of DM1 Mice.Mol Ther. 2019 Aug 7;27(8):1372-1388. doi: 10.1016/j.ymthe.2019.05.021. Epub 2019 Jun 5. Mol Ther. 2019. PMID: 31253581 Free PMC article.
-
Efficient CRISPR/Cas9-mediated editing of trinucleotide repeat expansion in myotonic dystrophy patient-derived iPS and myogenic cells.Nucleic Acids Res. 2018 Sep 19;46(16):8275-8298. doi: 10.1093/nar/gky548. Nucleic Acids Res. 2018. PMID: 29947794 Free PMC article.
-
Application of CRISPR-Cas9-Mediated Genome Editing for the Treatment of Myotonic Dystrophy Type 1.Mol Ther. 2020 Dec 2;28(12):2527-2539. doi: 10.1016/j.ymthe.2020.10.005. Epub 2020 Oct 14. Mol Ther. 2020. PMID: 33171139 Free PMC article. Review.
-
CRISPR/Cas Applications in Myotonic Dystrophy: Expanding Opportunities.Int J Mol Sci. 2019 Jul 27;20(15):3689. doi: 10.3390/ijms20153689. Int J Mol Sci. 2019. PMID: 31357652 Free PMC article. Review.
Cited by
-
Time-controlled and muscle-specific CRISPR/Cas9-mediated deletion of CTG-repeat expansion in the DMPK gene.Mol Ther Nucleic Acids. 2021 Nov 29;27:184-199. doi: 10.1016/j.omtn.2021.11.024. eCollection 2022 Mar 8. Mol Ther Nucleic Acids. 2021. PMID: 34976437 Free PMC article.
-
Therapeutic Genome Editing and In Vivo Delivery.AAPS J. 2021 Jun 2;23(4):80. doi: 10.1208/s12248-021-00613-w. AAPS J. 2021. PMID: 34080099 Review.
-
Evaluation of Engineered CRISPR-Cas-Mediated Systems for Site-Specific RNA Editing.Cell Rep. 2020 Nov 3;33(5):108350. doi: 10.1016/j.celrep.2020.108350. Cell Rep. 2020. PMID: 33147453 Free PMC article.
-
Therapeutic targeting of RNA for neurological and neuromuscular disease.Genes Dev. 2024 Sep 19;38(15-16):698-717. doi: 10.1101/gad.351612.124. Genes Dev. 2024. PMID: 39142832 Free PMC article. Review.
-
Detection of repeat expansions in large next generation DNA and RNA sequencing data without alignment.Sci Rep. 2022 Jul 30;12(1):13124. doi: 10.1038/s41598-022-17267-z. Sci Rep. 2022. PMID: 35907931 Free PMC article.
References
-
- Dion V Tissue specificity in DNA repair: lessons from trinucleotide repeat instability. Trends Genet. 30, 220–229 (2014). - PubMed
-
- Lopez Castel A, Cleary JD & Pearson CE Repeat instability as the basis for human diseases and as a potential target for therapy. Nat. Rev. Mol. Cell Biol. 11, 165–170 (2010). - PubMed
-
- Pearson CE Slipping while sleeping? Trinucleotide repeat expansions in germ cells. Trends Mol. Med. 9, 490–495 (2003). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

