Interaction Modes of Microsomal Cytochrome P450s with Its Reductase and the Role of Substrate Binding
- PMID: 32933097
- PMCID: PMC7555755
- DOI: 10.3390/ijms21186669
Interaction Modes of Microsomal Cytochrome P450s with Its Reductase and the Role of Substrate Binding
Abstract
The activity of microsomal cytochromes P450 (CYP) is strictly dependent on the supply of electrons provided by NADPH cytochrome P450 oxidoreductase (CPR). The variant nature of the isoform-specific proximal interface of microsomal CYPs implies that the interacting interface between the two proteins is degenerated. Recently, we demonstrated that specific CPR mutations in the FMN-domain (FD) may induce a gain in activity for a specific CYP isoform. In the current report, we confirm the CYP isoform dependence of CPR's degenerated binding by demonstrating that the effect of four of the formerly studied FD mutants are indeed exclusive of a specific CYP isoform, as verified by cytochrome c inhibition studies. Moreover, the nature of CYP's substrate seems to have a modulating role in the CPR:CYP interaction. In silico molecular dynamics simulations of the FD evidence that mutations induces very subtle structural alterations, influencing the characteristics of residues formerly implicated in the CPR:CYP interaction or in positioning of the FMN moiety. CPR seems therefore to be able to form effective interaction complexes with its structural diverse partners via a combination of specific structural features of the FD, which are functional in a CYP isoform dependent manner, and dependent on the substrate bound.
Keywords: CPR-FMN-domain; CYP-substrate; NADPH-cytochrome P450 reductase; electron transfer; microsomal cytochrome P450; protein–protein interaction.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
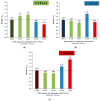





References
-
- Guengerich F.P. Human Cytochrome P450 Enzymes. In: Ortiz de Montellano P.R., editor. Cytochrome P450: Structure, Mechanism, and Biochemistry. 4th ed. Springer International Publishing; Cham, Switzerland: 2015. pp. 523–785.
-
- Strobel H.W., Hodgson A.V., Shen S. NADPH Cytochrome P450 Reductase and Its Structural and Functional Domains. In: Ortiz de Montellano P.R.O., editor. Cytochrome P450: Structure, Mechanism, and Biochemistry. 2nd ed. Springer; Boston, MA, USA: 1995. pp. 225–244.
-
- Waskell L., Kim J.-J.P. Electron Transfer Partners of Cytochrome P450. In: Ortiz de Montellano P.R., editor. Cytochrome P450: Structure, Mechanism, and Biochemistry. 4th ed. Springer International Publishing; Cham, Switzerland: 2015. pp. 33–68.
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

