A predictive index for health status using species-level gut microbiome profiling
- PMID: 32934239
- PMCID: PMC7492273
- DOI: 10.1038/s41467-020-18476-8
A predictive index for health status using species-level gut microbiome profiling
Abstract
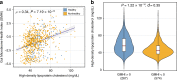
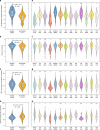
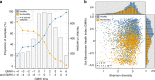
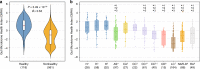
Providing insight into one's health status from a gut microbiome sample is an important clinical goal in current human microbiome research. Herein, we introduce the Gut Microbiome Health Index (GMHI), a biologically-interpretable mathematical formula for predicting the likelihood of disease independent of the clinical diagnosis. GMHI is formulated upon 50 microbial species associated with healthy gut ecosystems. These species are identified through a multi-study, integrative analysis on 4347 human stool metagenomes from 34 published studies across healthy and 12 different nonhealthy conditions, i.e., disease or abnormal bodyweight. When demonstrated on our population-scale meta-dataset, GMHI is the most robust and consistent predictor of disease presence (or absence) compared to α-diversity indices. Validation on 679 samples from 9 additional studies results in a balanced accuracy of 73.7% in distinguishing healthy from non-healthy groups. Our findings suggest that gut taxonomic signatures can predict health status, and highlight how data sharing efforts can provide broadly applicable discoveries.
Conflict of interest statement
V.K.G. and J.S. disclose that a patent application was filed relating to the materials in this manuscript. All other authors declare no competing interests.
Figures


References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

