Phosphoproteome Analysis in Immune Cell Signaling
- PMID: 32936995
- PMCID: PMC10961162
- DOI: 10.1002/cpim.105
Phosphoproteome Analysis in Immune Cell Signaling
Abstract
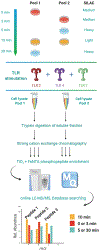
Immune cell signaling is largely regulated by protein phosphorylation. Stimulation of toll-like receptors (TLRs) by pathogen-associated ligands drives the cascade of immune response, which can be influenced by differences in phosphoprotein abundance. Therefore, the analysis of phosphorylation signatures at a global level is central to understanding the complex and integrated signaling in macrophages upon pathogen attack. Here, we describe a mass spectrometry-based approach to identify and quantify phosphoproteome changes in response to the stimulation of TLR2, TLR4, and TLR7 with immune-response inducing ligands in cultured immune cells. This review will focus on the TLR stimulation of mouse macrophages as an example; however, the technique is applicable to any immortalized immune cell and any soluble stimuli. The methodology includes protocols for metabolic labeling of immune cells (stable isotope labeling of amino acids in cell culture, i.e., SILAC); ligand-initiated stimulation of immune receptors followed by cell lysis; in-solution trypsin digestion of proteins and enrichment of the resulting peptide mix for collecting phosphopeptides, which are then analyzed by high-resolution LC-MS/MS (liquid-chromatography tandem mass spectrometry). © 2020 Wiley Periodicals LLC. Basic Protocol 1: SILAC labeling of mouse macrophages Basic Protocol 2: Stimulation, cell lysis and Western Blotting Basic Protocol 3: Trypsin digestion, fractionation and phosphopeptide enrichment Basic Protocol 4: Quantitative mass spectrometry Alternate Protocol: Culturing SILAC-labeled cells from frozen mouse macrophages cells.
Keywords: cell signaling; innate immunity; mass spectrometry; phosphoproteomics.
© 2020 Wiley Periodicals LLC.
Figures



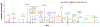
Similar articles
-
A Methodology for Comprehensive Analysis of Toll-Like Receptor Signaling in Macrophages.Methods Mol Biol. 2017;1636:301-312. doi: 10.1007/978-1-4939-7154-1_19. Methods Mol Biol. 2017. PMID: 28730487 Free PMC article.
-
Phosphoproteome profiling of the macrophage response to different toll-like receptor ligands identifies differences in global phosphorylation dynamics.J Proteome Res. 2014 Nov 7;13(11):5185-97. doi: 10.1021/pr5002466. Epub 2014 Jun 18. J Proteome Res. 2014. PMID: 24941444 Free PMC article.
-
Fractionation of Enriched Phosphopeptides Using pH/Acetonitrile-Gradient-Reversed-Phase Microcolumn Separation in Combination with LC-MS/MS Analysis.Int J Mol Sci. 2020 Jun 1;21(11):3971. doi: 10.3390/ijms21113971. Int J Mol Sci. 2020. PMID: 32492839 Free PMC article.
-
Phosphoproteomic analysis: an emerging role in deciphering cellular signaling in human embryonic stem cells and their differentiated derivatives.Stem Cell Rev Rep. 2012 Mar;8(1):16-31. doi: 10.1007/s12015-011-9317-8. Stem Cell Rev Rep. 2012. PMID: 22009073 Free PMC article. Review.
-
Quantitative analysis of global phosphorylation changes with high-resolution tandem mass spectrometry and stable isotopic labeling.Methods. 2013 Jun 15;61(3):251-9. doi: 10.1016/j.ymeth.2013.04.010. Epub 2013 Apr 21. Methods. 2013. PMID: 23611819 Free PMC article. Review.
Cited by
-
Interrogating Host Antiviral Environments Driven by Nuclear DNA Sensing: A Multiomic Perspective.Biomolecules. 2020 Nov 24;10(12):1591. doi: 10.3390/biom10121591. Biomolecules. 2020. PMID: 33255247 Free PMC article. Review.
-
Association of TLR4 gene 2026A/G (rs1927914), 896A/G (rs4986790), and 1196C/T (rs4986791) polymorphisms and cancer susceptibility: Meta-analysis and trial sequential analysis.Medicine (Baltimore). 2023 Feb 22;102(8):e33040. doi: 10.1097/MD.0000000000033040. Medicine (Baltimore). 2023. PMID: 36827055 Free PMC article.
-
Usp9x contributes to the development of sepsis-induced acute kidney injury by promoting inflammation and apoptosis in renal tubular epithelial cells via activation of the TLR4/nf-κb pathway.Ren Fail. 2024 Dec;46(2):2361089. doi: 10.1080/0886022X.2024.2361089. Epub 2024 Jun 14. Ren Fail. 2024. PMID: 38874156 Free PMC article.
References
Literature Cited
-
- Du R, Long J, Yao J, Dong Y, Yang X, Tang S, … Chen X (2010). Subcellular quantitative proteomics reveals multiple pathway crosstalk that coordinates specific signaling and transcriptional regulation for the early host response to LPS. Journal of Proteome Research, 9(4), 1805–1821. doi: 10.1021/pr900962c. - DOI - PubMed
-
- Hoedt E, Zhang G, & Neubert TA (2019). Stable isotope labeling by amino acids in cell culture (SILAC) for quantitative proteomics. In Woods AG & Darie CC (Eds.), Advancements of mass spectrometry in biomedical research (pp. 531–539). Cham: Springer International Publishing.
Internet Resource
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

