Databases and Bioinformatic Tools for Glycobiology and Glycoproteomics
- PMID: 32937895
- PMCID: PMC7556027
- DOI: 10.3390/ijms21186727
Databases and Bioinformatic Tools for Glycobiology and Glycoproteomics
Abstract
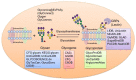
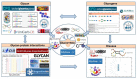
Glycosylation plays critical roles in various biological processes and is closely related to diseases. Deciphering the glycocode in diverse cells and tissues offers opportunities to develop new disease biomarkers and more effective recombinant therapeutics. In the past few decades, with the development of glycobiology, glycomics, and glycoproteomics technologies, a large amount of glycoscience data has been generated. Subsequently, a number of glycobiology databases covering glycan structure, the glycosylation sites, the protein scaffolds, and related glycogenes have been developed to store, analyze, and integrate these data. However, these databases and tools are not well known or widely used by the public, including clinicians and other researchers who are not in the field of glycobiology, but are interested in glycoproteins. In this study, the representative databases of glycan structure, glycoprotein, glycan-protein interactions, glycogenes, and the newly developed bioinformatic tools and integrated portal for glycoproteomics are reviewed. We hope this overview could assist readers in searching for information on glycoproteins of interest, and promote further clinical application of glycobiology.
Keywords: glycan; glycogene; glycoinformatics; glycoprotein; glycosylation; lectin.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



References
-
- Strum J.S., Nwosu C.C., Hua S., Kronewitter S.R., Seipert R.R., Bachelor R.J., An H.J., Lebrilla C.B. Automated assignments of n- and o-site specific glycosylation with extensive glycan heterogeneity of glycoprotein mixtures. Anal. Chem. 2013;85:5666–5675. doi: 10.1021/ac4006556. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

