Chromatin remodeling in bovine embryos indicates species-specific regulation of genome activation
- PMID: 32943640
- PMCID: PMC7498599
- DOI: 10.1038/s41467-020-18508-3
Chromatin remodeling in bovine embryos indicates species-specific regulation of genome activation
Abstract
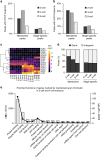
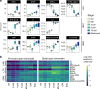
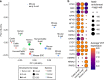
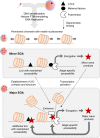
The shift from maternal to embryonic control is a critical developmental milestone in preimplantation development. Widespread transcriptomic and epigenetic remodeling facilitate this transition from terminally differentiated gametes to totipotent blastomeres, but the identity of transcription factors (TF) and genomic elements regulating embryonic genome activation (EGA) are poorly defined. The timing of EGA is species-specific, e.g., the timing of murine and human EGA differ significantly. To deepen our understanding of mammalian EGA, here we profile changes in open chromatin during bovine preimplantation development. Before EGA, open chromatin is enriched for maternal TF binding, similar to that observed in humans and mice. During EGA, homeobox factor binding becomes more prevalent and requires embryonic transcription. A cross-species comparison of open chromatin during preimplantation development reveals strong similarity in the regulatory circuitry underlying bovine and human EGA compared to mouse. Moreover, TFs associated with murine EGA are not enriched in cattle or humans, indicating that cattle may be a more informative model for human preimplantation development than mice.
Conflict of interest statement
The authors declare no competing interests.
Figures





Similar articles
-
Analysis of active chromatin modifications in early mammalian embryos reveals uncoupling of H2A.Z acetylation and H3K36 trimethylation from embryonic genome activation.Epigenetics. 2012 Jul;7(7):747-57. doi: 10.4161/epi.20584. Epub 2012 Jul 1. Epigenetics. 2012. PMID: 22647320
-
Molecular cloning of PRD-like homeobox genes expressed in bovine oocytes and early IVF embryos.BMC Genomics. 2024 Nov 6;25(1):1048. doi: 10.1186/s12864-024-10969-w. BMC Genomics. 2024. PMID: 39506635 Free PMC article.
-
Genome activation in bovine embryos: review of the literature and new insights from RNA sequencing experiments.Anim Reprod Sci. 2014 Sep;149(1-2):46-58. doi: 10.1016/j.anireprosci.2014.05.016. Epub 2014 Jun 6. Anim Reprod Sci. 2014. PMID: 24975847 Review.
-
Functional genomics of HMGN3a and SMARCAL1 in early mammalian embryogenesis.BMC Genomics. 2009 Apr 24;10:183. doi: 10.1186/1471-2164-10-183. BMC Genomics. 2009. PMID: 19393058 Free PMC article.
-
The initiation of mammalian embryonic transcription: to begin at the beginning.Trends Cell Biol. 2023 May;33(5):365-373. doi: 10.1016/j.tcb.2022.08.008. Epub 2022 Sep 29. Trends Cell Biol. 2023. PMID: 36182534 Review.
Cited by
-
Evaluating histone modification analysis of individual preimplantation embryos.BMC Genomics. 2024 Jan 18;25(1):75. doi: 10.1186/s12864-024-09984-8. BMC Genomics. 2024. PMID: 38238676 Free PMC article.
-
Profiling of open chromatin in developing pig (Sus scrofa) muscle to identify regulatory regions.G3 (Bethesda). 2022 Feb 4;12(2):jkab424. doi: 10.1093/g3journal/jkab424. G3 (Bethesda). 2022. PMID: 34897420 Free PMC article.
-
Multiple Retrotransposon-mediated NF-YA Gene Duplication Events Recurred in Diverse Groups of Mammals at Different Ancestry Levels.Genome Biol Evol. 2025 Apr 30;17(5):evaf071. doi: 10.1093/gbe/evaf071. Genome Biol Evol. 2025. PMID: 40408078 Free PMC article.
-
Dynamic Change of R-Loop Implicates in the Regulation of Zygotic Genome Activation in Mouse.Int J Mol Sci. 2022 Nov 18;23(22):14345. doi: 10.3390/ijms232214345. Int J Mol Sci. 2022. PMID: 36430821 Free PMC article.
-
The Epigenetics of Gametes and Early Embryos and Potential Long-Range Consequences in Livestock Species-Filling in the Picture With Epigenomic Analyses.Front Genet. 2021 Mar 3;12:557934. doi: 10.3389/fgene.2021.557934. eCollection 2021. Front Genet. 2021. PMID: 33747031 Free PMC article. Review.
References
-
- Ram PT, Schultz RM. Reporter gene expression in G2 of the 1-cell mouse embryo. Dev. Biol. 1993;156:552–556. - PubMed
-
- Latham KE, Garrels JI, Chang C, Solter D. Quantitative analysis of protein synthesis in mouse embryos. I. Extensive reprogramming at the one- and two-cell stages. Development. 1991;112:921 LP–921932. - PubMed
-
- Eckersley-Maslin MA, Alda-Catalinas C, Reik W. Dynamics of the epigenetic landscape during the maternal-to-zygotic transition. Nat. Rev. Mol. Cell Biol. 2018;19:436–450. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

