Parvimonas micra as a putative non-invasive faecal biomarker for colorectal cancer
- PMID: 32943695
- PMCID: PMC7499209
- DOI: 10.1038/s41598-020-72132-1
Parvimonas micra as a putative non-invasive faecal biomarker for colorectal cancer
Abstract
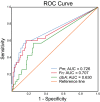
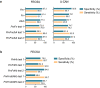
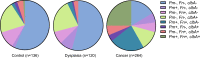
The use of faecal microbial markers as non-invasive biomarkers for colorectal cancer (CRC) has been suggested, but not fully elucidated. Here, we have evaluated the importance of Parvimonas micra as a potential non-invasive faecal biomarker in CRC and its relation to other microbial biomarkers. The levels of P. micra, F. nucleatum and clbA + bacteria were quantified using qPCR in faecal samples from a population-based cohort of patients undergoing colonoscopy due to symptoms from the large bowel. The study included 38 CRC patients, 128 patients with dysplasia and 63 controls. The results were validated in a second consecutive CRC cohort including faecal samples from 238 CRC patients and 94 controls. We found significantly higher levels of P. micra in faecal samples from CRC patients compared to controls. A test for P. micra could detect CRC with a specificity of 87.3% and a sensitivity of 60.5%. In addition, we found that combining P. micra with other microbial markers, could further enhance test sensitivity. Our findings support the potential use of P. micra as a non-invasive biomarker for CRC. Together with other microbial faecal markers, P. micra may identify patients with "high risk" microbial patterns, indicating increased risk and incidence of cancer.
Conflict of interest statement
The authors declare no competing interests.
Figures

Similar articles
-
Metagenomic analysis of faecal microbiome as a tool towards targeted non-invasive biomarkers for colorectal cancer.Gut. 2017 Jan;66(1):70-78. doi: 10.1136/gutjnl-2015-309800. Epub 2015 Sep 25. Gut. 2017. PMID: 26408641
-
Parvimonas micra, Peptostreptococcus stomatis, Fusobacterium nucleatum and Akkermansia muciniphila as a four-bacteria biomarker panel of colorectal cancer.Sci Rep. 2021 Feb 3;11(1):2925. doi: 10.1038/s41598-021-82465-0. Sci Rep. 2021. PMID: 33536501 Free PMC article.
-
Parvimonas micra forms a distinct bacterial network with oral pathobionts in colorectal cancer patients.J Transl Med. 2024 Oct 17;22(1):947. doi: 10.1186/s12967-024-05720-8. J Transl Med. 2024. PMID: 39420333 Free PMC article.
-
Faecal microbial biomarkers in early diagnosis of colorectal cancer.J Cell Mol Med. 2021 Dec;25(23):10783-10797. doi: 10.1111/jcmm.17010. Epub 2021 Nov 9. J Cell Mol Med. 2021. PMID: 34750964 Free PMC article. Review.
-
Microbial markers in colorectal cancer detection and/or prognosis.World J Gastroenterol. 2018 Jun 14;24(22):2327-2347. doi: 10.3748/wjg.v24.i22.2327. World J Gastroenterol. 2018. PMID: 29904241 Free PMC article. Review.
Cited by
-
Characterization of Lung and Oral Microbiomes in Lung Cancer Patients Using Culturomics and 16S rRNA Gene Sequencing.Microbiol Spectr. 2023 Jun 15;11(3):e0031423. doi: 10.1128/spectrum.00314-23. Epub 2023 Apr 24. Microbiol Spectr. 2023. PMID: 37092999 Free PMC article.
-
Detection of colorectal-cancer-associated bacterial taxa in fecal samples using next-generation sequencing and 19 newly established qPCR assays.Mol Oncol. 2025 Feb;19(2):412-429. doi: 10.1002/1878-0261.13700. Epub 2024 Jul 6. Mol Oncol. 2025. PMID: 38970464 Free PMC article.
-
An association between poor oral health, oral microbiota, and pain identified in New Zealand women with central sensitisation disorders: a prospective clinical study.Front Pain Res (Lausanne). 2025 Apr 9;6:1577193. doi: 10.3389/fpain.2025.1577193. eCollection 2025. Front Pain Res (Lausanne). 2025. PMID: 40270934 Free PMC article.
-
Gut Microbiota in Colorectal Cancer: Biological Role and Therapeutic Opportunities.Cancers (Basel). 2023 Jan 30;15(3):866. doi: 10.3390/cancers15030866. Cancers (Basel). 2023. PMID: 36765824 Free PMC article. Review.
-
Gut Microbiota: A Potential Target for Cancer Interventions.Cancer Manag Res. 2021 Nov 3;13:8281-8296. doi: 10.2147/CMAR.S328249. eCollection 2021. Cancer Manag Res. 2021. PMID: 34764691 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

