Revealing antimicrobial resistance in stormwater with MinION
- PMID: 32947654
- PMCID: PMC7297696
- DOI: 10.1016/j.chemosphere.2020.127392
Revealing antimicrobial resistance in stormwater with MinION
Abstract
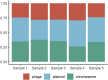
Discharge of urban stormwater containing organic matter, heavy metals and sometime human feces, to the natural aquatic reservoirs without any treatment is not only an environmental problem. It can lead to prevalence of antibiotic resistant bacteria in stormwater systems and transmission of antibiotic resistance genes to the environment. We performed antibiotic resistome identification and virus detection in stormwater samples from Stockholm, using publicly available metagenomic sequencing MinION data. A MinION platform offers low-cost, precise environmental metagenomics analysis. 37 groups of antibiotic resistant bacteria (ARB), 11 resistance types with 26 resistance mechanisms - antibiotic resistance genes (ARGs) giving tolerance to the aminoglycoside, beta-lactams, fosmidomycin, MLS, multidrug and vancomycin were identified using ARGpore pipeline. The majority of the identified bacteria species were related to the natural environment such as soil and were not dangerous to human. Alarmingly, human pathogenic bacteria carrying resistance to antibiotics currently used against them (Bordetella resistant to macrolides and multidrug resistant Propionibacterium avidum) were also found in the samples. Most abundant viruses identified belonged to Caudovirales and Herpesvirales and they were not carrying ARGs. Unlike the virome, resistome and ARB were not unique for stormwater sampling points. This results underline the need for extensive monitoring of the microbial community structure in the urban stormwater systems to assess antimicrobial resistance spread.
Keywords: ARB; ARG; Metagenomics; MinION; Stormwater.
Copyright © 2020 The Authors. Published by Elsevier Ltd.. All rights reserved.
Conflict of interest statement
Declaration of competing interest The authors, Maciej Białasek and Aleksandra Miłobędzka declare no conflict of interest.
Figures


Similar articles
-
Metagenomic profiling of raw wastewater in Portugal highlights microbiota and resistome signatures of public health interest beyond the usual suspects.Sci Total Environ. 2024 Oct 10;946:174272. doi: 10.1016/j.scitotenv.2024.174272. Epub 2024 Jun 24. Sci Total Environ. 2024. PMID: 38925382
-
Bioprospecting for β-lactam resistance genes using a metagenomics-guided strategy.Appl Microbiol Biotechnol. 2017 Aug;101(15):6253-6260. doi: 10.1007/s00253-017-8343-0. Epub 2017 Jun 5. Appl Microbiol Biotechnol. 2017. PMID: 28584911
-
Metagenomic Assembly Reveals Hosts of Antibiotic Resistance Genes and the Shared Resistome in Pig, Chicken, and Human Feces.Environ Sci Technol. 2016 Jan 5;50(1):420-7. doi: 10.1021/acs.est.5b03522. Epub 2015 Dec 22. Environ Sci Technol. 2016. PMID: 26650334
-
Urban wastewater treatment plants as hotspots for antibiotic resistant bacteria and genes spread into the environment: a review.Sci Total Environ. 2013 Mar 1;447:345-60. doi: 10.1016/j.scitotenv.2013.01.032. Epub 2013 Feb 7. Sci Total Environ. 2013. PMID: 23396083 Review.
-
Functional Metagenomics as a Tool for Identification of New Antibiotic Resistance Genes from Natural Environments.Microb Ecol. 2017 Feb;73(2):479-491. doi: 10.1007/s00248-016-0866-x. Epub 2016 Oct 5. Microb Ecol. 2017. PMID: 27709246 Review.
Cited by
-
A resistome survey across hundreds of freshwater bacterial communities reveals the impacts of veterinary and human antibiotics use.Front Microbiol. 2022 Oct 6;13:995418. doi: 10.3389/fmicb.2022.995418. eCollection 2022. Front Microbiol. 2022. PMID: 36338036 Free PMC article.
-
A review of the resistome within the digestive tract of livestock.J Anim Sci Biotechnol. 2021 Nov 11;12(1):121. doi: 10.1186/s40104-021-00643-6. J Anim Sci Biotechnol. 2021. PMID: 34763729 Free PMC article. Review.
-
Implementing Wastewater-Based Epidemiology for Long-Read Metagenomic Sequencing of Antimicrobial Resistance in Kampala, Uganda.Microorganisms. 2025 May 28;13(6):1240. doi: 10.3390/microorganisms13061240. Microorganisms. 2025. PMID: 40572128 Free PMC article.
References
-
- Alonso A., Rojo F., Martínez J.L. Environmental and clinical isolates of Pseudomonas aeruginosa show pathogenic and biodegradative properties irrespective of their origin. Environ. Microbiol. 1999;1:421–430. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

