Network and Systems Medicine: Position Paper of the European Collaboration on Science and Technology Action on Open Multiscale Systems Medicine
- PMID: 32954378
- PMCID: PMC7500076
- DOI: 10.1089/nsm.2020.0004
Network and Systems Medicine: Position Paper of the European Collaboration on Science and Technology Action on Open Multiscale Systems Medicine
Abstract
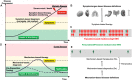
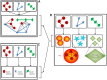
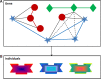
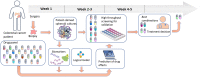
Introduction: Network and systems medicine has rapidly evolved over the past decade, thanks to computational and integrative tools, which stem in part from systems biology. However, major challenges and hurdles are still present regarding validation and translation into clinical application and decision making for precision medicine. Methods: In this context, the Collaboration on Science and Technology Action on Open Multiscale Systems Medicine (OpenMultiMed) reviewed the available advanced technologies for multidimensional data generation and integration in an open-science approach as well as key clinical applications of network and systems medicine and the main issues and opportunities for the future. Results: The development of multi-omic approaches as well as new digital tools provides a unique opportunity to explore complex biological systems and networks at different scales. Moreover, the application of findable, applicable, interoperable, and reusable principles and the adoption of standards increases data availability and sharing for multiscale integration and interpretation. These innovations have led to the first clinical applications of network and systems medicine, particularly in the field of personalized therapy and drug dosing. Enlarging network and systems medicine application would now imply to increase patient engagement and health care providers as well as to educate the novel generations of medical doctors and biomedical researchers to shift the current organ- and symptom-based medical concepts toward network- and systems-based ones for more precise diagnoses, interventions, and ideally prevention. Conclusion: In this dynamic setting, the health care system will also have to evolve, if not revolutionize, in terms of organization and management.
Keywords: big data; data integration; integrated health care; omics; systems medicine.
© Blandine Comte et al. 2020; Published by Mary Ann Liebert, Inc.
Conflict of interest statement
No competing financial interests exist.
Figures
References
-
- Nosengo N. Can you teach old drugs new tricks? Nature. 2016;534:314–316 - PubMed
-
- Wieseler B, McGauran N, Kaiser T. New drugs: where did we go wrong and what can we do better? BMJ. 2019;l4340. - PubMed
-
- Schork NJ. Personalized medicine: time for one-person trials. Nature. 2015;520:609–611 - PubMed
-
- Scannell JW, Blanckley A, Boldon H, et al. . Diagnosing the decline in pharmaceutical R&D efficiency. Nat Rev Drug Discov. 2012;11:191–200 - PubMed
Publication types
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
