Comparison of three commercial decision support platforms for matching of next-generation sequencing results with therapies in patients with cancer
- PMID: 32967919
- PMCID: PMC7513637
- DOI: 10.1136/esmoopen-2020-000872
Comparison of three commercial decision support platforms for matching of next-generation sequencing results with therapies in patients with cancer
Abstract
Objective: Precision oncology depends on translating molecular data into therapy recommendations. However, with the growing complexity of next-generation sequencing-based tests, clinical interpretation of somatic genomic mutations has evolved into a formidable task. Here, we compared the performance of three commercial clinical decision support tools, that is, NAVIFY Mutation Profiler (NAVIFY; Roche), QIAGEN Clinical Insight (QCI) Interpret (QIAGEN) and CureMatch Bionov (CureMatch).
Methods: In order to obtain the current status of the respective tumour genome, we analysed cell-free DNA from patients with metastatic breast, colorectal or non-small cell lung cancer. We evaluated somatic copy number alterations and in parallel applied a 77-gene panel (AVENIO ctDNA Expanded Panel). We then assessed the concordance of tier classification approaches between NAVIFY and QCI and compared the strategies to determine actionability among all three platforms. Finally, we quantified the alignment of treatment suggestions across all decision tools.
Results: Each platform varied in its mode of variant classification and strategy for identifying druggable targets and clinical trials, which resulted in major discrepancies. Even the frequency of concordant actionable events for tier I-A or tier I-B classifications was only 4.3%, 9.5% and 28.4% when comparing NAVIFY with QCI, NAVIFY with CureMatch and CureMatch with QCI, respectively, and the obtained treatment recommendations differed drastically.
Conclusions: Treatment decisions based on molecular markers appear at present to be arbitrary and dependent on the chosen strategy. As a consequence, tumours with identical molecular profiles would be differently treated, which challenges the promising concepts of genome-informed medicine.
Keywords: circulating tumour DNA; clinical decision support; molecular profiling; next-generation sequencing; variant interpretation.
© Author (s) (or their employer(s)) 2020. Re-use permitted under CC BY-NC. No commercial re-use. Published by BMJ on behalf of the European Society for Medical Oncology.
Conflict of interest statement
Competing interests: EH and MRS have an unrelated sponsored research agreement with Servier within CANCER-ID, a project funded by the Innovative Medicines Joint Undertaking (IMI JU). EH receives funding from Freenome, South San Francisco, CA and PreAnalytiX, Hombrechtikon, Switzerland. Roche Diagnostics and QIAGEN provided the authors with free access to their respective platforms for a restricted time period to facilitate this study. EH has received honoraria for advisory boards from Roche. ES has served as consultant or on the advisory board for AstraZeneca, Roche, Pfizer, Novartis, MSD/Merck, Bayer and Janssen Cilag (Johnson&Johnson). The other authors have no competing interests to declare.
Figures

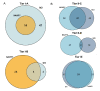
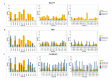

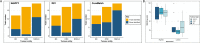
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

