Initiation of a conserved trophectoderm program in human, cow and mouse embryos
- PMID: 32968278
- PMCID: PMC7116563
- DOI: 10.1038/s41586-020-2759-x
Initiation of a conserved trophectoderm program in human, cow and mouse embryos
Abstract
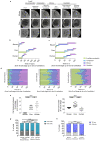
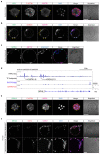
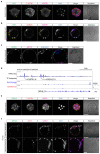
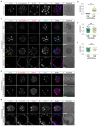
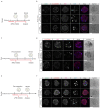
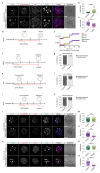
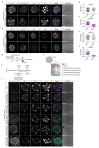
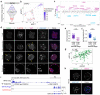
Current understandings of cell specification in early mammalian pre-implantation development are based mainly on mouse studies. The first lineage differentiation event occurs at the morula stage, with outer cells initiating a trophectoderm (TE) placental progenitor program. The inner cell mass arises from inner cells during subsequent developmental stages and comprises precursor cells of the embryo proper and yolk sac1. Recent gene-expression analyses suggest that the mechanisms that regulate early lineage specification in the mouse may differ in other mammals, including human2-5 and cow6. Here we show the evolutionary conservation of a molecular cascade that initiates TE segregation in human, cow and mouse embryos. At the morula stage, outer cells acquire an apical-basal cell polarity, with expression of atypical protein kinase C (aPKC) at the contact-free domain, nuclear expression of Hippo signalling pathway effectors and restricted expression of TE-associated factors such as GATA3, which suggests initiation of a TE program. Furthermore, we demonstrate that inhibition of aPKC by small-molecule pharmacological modulation or Trim-Away protein depletion impairs TE initiation at the morula stage. Our comparative embryology analysis provides insights into early lineage specification and suggests that a similar mechanism initiates a TE program in human, cow and mouse embryos.
Conflict of interest statement
All authors declare no competing financial interests.
Figures





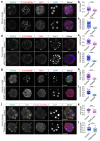
Comment in
-
Universal assembly instructions for the placenta.Nature. 2020 Nov;587(7834):370-371. doi: 10.1038/d41586-020-02914-0. Nature. 2020. PMID: 33067590 No abstract available.
References
-
- Fogarty NME, et al. Genome editing reveals a role for OCT4 in human embryogenesis. Nature. 2017;550:67. doi: 10.1038/nature24033. https://www.nature.com/articles/nature24033#supplementary-information. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

