Bone morphogenetic protein 7 promotes resistance to immunotherapy
- PMID: 32973129
- PMCID: PMC7519103
- DOI: 10.1038/s41467-020-18617-z
Bone morphogenetic protein 7 promotes resistance to immunotherapy
Erratum in
-
Author Correction: Bone morphogenetic protein 7 promotes resistance to immunotherapy.Nat Commun. 2020 Oct 8;11(1):5144. doi: 10.1038/s41467-020-19083-3. Nat Commun. 2020. PMID: 33033261 Free PMC article.
Abstract
Immunotherapies revolutionized cancer treatment by harnessing the immune system to target cancer cells. However, most patients are resistant to immunotherapies and the mechanisms underlying this resistant is still poorly understood. Here, we report that overexpression of BMP7, a member of the TGFB superfamily, represents a mechanism for resistance to anti-PD1 therapy in preclinical models and in patients with disease progression while on immunotherapies. BMP7 secreted by tumor cells acts on macrophages and CD4+ T cells in the tumor microenvironment, inhibiting MAPK14 expression and impairing pro-inflammatory responses. Knockdown of BMP7 or its neutralization via follistatin in combination with anti-PD1 re-sensitizes resistant tumors to immunotherapies. Thus, we identify the BMP7 signaling pathway as a potential immunotherapeutic target in cancer.
Conflict of interest statement
J.W.W. reports research support from GlaxoSmithKline, Bristol Meyers Squibb, Merck, Nanobiotix, Mavu Pharma, Takeda, Varian, and Checkmate Pharmaceuticals. J.W.W. serves on the scientific advisory board for Legion Healthcare Partners, RefleXion Medical, MolecularMatch, Merck, AstraZeneca, Aileron Therapeutics, Ventana, OncoResponse, CheckMate Pharmaceuticals, Mavu Pharma, Alpine Immune Sciences, and Nanorobotix. He is co-founder of Healios, MolecularMatch, and OncoResponse and serves as an advisor to Astra Zeneca, Merck, MolecularMatch, Incyte, Aileron, and Nanobiotix. J.W.W. holds stock or ownership in Alpine Immune Sciences, Checkmate Pharmaceuticals, Healios, Mavu Pharma, Legion Healthcare Partners, MolecularMatch, Nanorobotix, OncoResponse, and RefleXion. J.W.W. has accepted honoraria in the form of travel costs from Nanobiotix, RefleXion, Varian, Shandong University, The Korea Society of Radiology, Aileron Therapeutics and Ventana. J.W.W. has the following patents; MP470 (amuvatinib), MRX34 regulation of PDL1, XRT technique to overcome immune resistance. MD Anderson Cancer Center has a trademark for RadScopalTM. P.H. receives laboratory research support from Dragonfly, Immatics, Sanofi, and GlaxoSmithKline. All other authors declare no conflicts of interest.
Figures


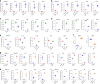
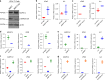

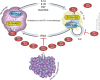
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

