Multiple Processes May Involve in the IgG4-RD Pathogenesis: An Integrative Study via Proteomic and Transcriptomic Analysis
- PMID: 32973752
- PMCID: PMC7468437
- DOI: 10.3389/fimmu.2020.01795
Multiple Processes May Involve in the IgG4-RD Pathogenesis: An Integrative Study via Proteomic and Transcriptomic Analysis
Abstract
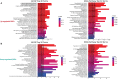
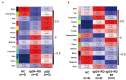
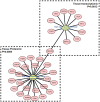
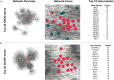
Immunoglobulin G4-related disease (IgG4-RD) is a newly defined disease entity, while the exact pathogenesis is still not clear. Identifying the characters of IgG4-RD in proteomic and transcriptomic aspects will be critical to investigate the potential pathogenic mechanisms of IgG4-RD. We performed proteomic analysis realized with iTRAQ technique for serum samples from eight treatment-naive IgG4-RD patients and eight healthy volunteers, and tissue samples from two IgG4-RD patients and two non-IgG4-RD patients. Transcriptomic data (GSE40568 and GSE66465) was obtained from the GEO Dataset for validation. The weighted correlation network analysis (WGCNA) was applied to detect the gene modules correlated with IgG4-RD. KEGG pathway analysis was used to investigate pathways enriched in IgG4-RD samples. As a result, a total of 980 differentially expressed proteins (DEPs) in tissue and 94 DEPs in serum were identified between IgG4-RD and control groups. Three hundred fifty-four and two hundred forty-seven genes that most correlated with IgG4-RD were detected by WGCNA analysis in tissue and PBMC, respectively. We also found that DEPs in IgG4-RD samples were enriched in several immune-related activities including bacterial/viral infections and platelet activation as well as some immune related signaling pathways. In conclusion, we identified multiple processes/factors and several signaling pathways that may involve in the IgG4-RD pathogenesis, and found out some potential therapeutic targets for IgG4-RD.
Keywords: IgG4-RD pathogenesis; IgG4-related disease (IgG4-RD); WGCNA (Weighted Gene Co-expression Network Analyses); enrichment analysis; proteomic analysis.
Copyright © 2020 Cai, Chen, Lin, Ye, Zheng and Dong.
Figures



References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

