Alginate genes are required for optimal soil colonization and persistence by Pseudomonas fluorescens Pf0-1
- PMID: 32974516
- PMCID: PMC7471777
- DOI: 10.1099/acmi.0.000021
Alginate genes are required for optimal soil colonization and persistence by Pseudomonas fluorescens Pf0-1
Abstract
Pseudomonas fluorescens strains are important candidates for use as biological control agents to reduce fungal diseases on crop plants. To understand the ecological success of these bacteria and for successful and stable biological control, determination of how these bacteria colonize and persist in soil environments is critical. Here we show that P. fluorescens Pf0-1 is negatively impacted by reduced water availability in soil, but adapts and persists. A pilot transcriptomic study of Pf0-1 colonizing moist and dehydrated soil was used to identify candidate genetic loci, which could play a role in the adaptation to dehydration. Genes predicted to specify alginate production were identified and chosen for functional evaluation. Using deletion mutants, predicted alginate biosynthesis genes were shown to be important for optimal colonization of moist soil, and necessary for adaptation to reduced water availability in dried soil. Our findings extend in vitro studies of water stress into a more natural system and suggest alginate may be an essential extracellular product for the lifestyle of P. fluorescens when growing in soil.
Keywords: Pseudomonas fluorescens; biological control; soil colonization; water stress.
© 2019 The Authors.
Conflict of interest statement
The authors declare that there are no conflicts of interest.
Figures

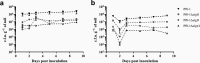
Similar articles
-
Colonization strategies of Pseudomonas fluorescens Pf0-1: activation of soil-specific genes important for diverse and specific environments.BMC Microbiol. 2013 Apr 27;13:92. doi: 10.1186/1471-2180-13-92. BMC Microbiol. 2013. PMID: 23622502 Free PMC article.
-
Requirement of polyphosphate by Pseudomonas fluorescens Pf0-1 for competitive fitness and heat tolerance in laboratory media and sterile soil.Appl Environ Microbiol. 2009 Jun;75(12):3872-81. doi: 10.1128/AEM.00017-09. Epub 2009 Apr 24. Appl Environ Microbiol. 2009. PMID: 19395572 Free PMC article.
-
Overlapping protein-encoding genes in Pseudomonas fluorescens Pf0-1.PLoS Genet. 2008 Jun 13;4(6):e1000094. doi: 10.1371/journal.pgen.1000094. PLoS Genet. 2008. PMID: 18551168 Free PMC article.
-
The adnA transcriptional factor affects persistence and spread of Pseudomonas fluorescens under natural field conditions.Appl Environ Microbiol. 2001 Feb;67(2):852-7. doi: 10.1128/AEM.67.2.852-857.2001. Appl Environ Microbiol. 2001. PMID: 11157254 Free PMC article.
-
New insights into Pseudomonas fluorescens alginate biosynthesis relevant for the establishment of an efficient production process for microbial alginates.N Biotechnol. 2017 Jul 25;37(Pt A):2-8. doi: 10.1016/j.nbt.2016.08.005. Epub 2016 Sep 1. N Biotechnol. 2017. PMID: 27593394 Review.
Cited by
-
Leveraging Pseudomonas Stress Response Mechanisms for Industrial Applications.Front Microbiol. 2021 May 10;12:660134. doi: 10.3389/fmicb.2021.660134. eCollection 2021. Front Microbiol. 2021. PMID: 34040596 Free PMC article. Review.
-
PelD is required downstream of c-di-GMP for host specialization of Pseudomonas lurida.BMC Microbiol. 2025 Apr 16;25(1):220. doi: 10.1186/s12866-025-03945-1. BMC Microbiol. 2025. PMID: 40241006 Free PMC article.
-
Genetic factors involved in rhizosphere colonization by phytobeneficial Pseudomonas spp.Comput Struct Biotechnol J. 2020 Nov 19;18:3539-3554. doi: 10.1016/j.csbj.2020.11.025. eCollection 2020. Comput Struct Biotechnol J. 2020. PMID: 33304453 Free PMC article. Review.
-
Corner Flows Induced by Surfactant-Producing Bacteria Bacillus subtilis and Pseudomonas fluorescens.Microbiol Spectr. 2022 Oct 26;10(5):e0323322. doi: 10.1128/spectrum.03233-22. Epub 2022 Oct 10. Microbiol Spectr. 2022. PMID: 36214703 Free PMC article.
-
Transcriptional Response and Plant Growth Promoting Activity of Pseudomonas fluorescens DR397 under Drought Stress Conditions.Microbiol Spectr. 2022 Aug 31;10(4):e0097922. doi: 10.1128/spectrum.00979-22. Epub 2022 Jul 12. Microbiol Spectr. 2022. PMID: 35863006 Free PMC article.
References
-
- Loper JE, Gross H. Genomic analysis of antifungal metabolite production by Pseudomonas fluorescens Pf-5. Eur J Plant Pathol. 2007;119:265–278. doi: 10.1007/s10658-007-9179-8. - DOI
LinkOut - more resources
Full Text Sources
