TREND-DB-a transcriptome-wide atlas of the dynamic landscape of alternative polyadenylation
- PMID: 32976578
- PMCID: PMC7778938
- DOI: 10.1093/nar/gkaa722
TREND-DB-a transcriptome-wide atlas of the dynamic landscape of alternative polyadenylation
Abstract
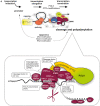
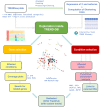
Alternative polyadenylation (APA) profoundly expands the transcriptome complexity. Perturbations of APA can disrupt biological processes, ultimately resulting in devastating disorders. A major challenge in identifying mechanisms and consequences of APA (and its perturbations) lies in the complexity of RNA 3' end processing, involving poorly conserved RNA motifs and multi-component complexes consisting of far more than 50 proteins. This is further complicated in that RNA 3' end maturation is closely linked to transcription, RNA processing and even epigenetic (histone/DNA/RNA) modifications. Here, we present TREND-DB (http://shiny.imbei.uni-mainz.de:3838/trend-db), a resource cataloging the dynamic landscape of APA after depletion of >170 proteins involved in various facets of transcriptional, co- and post-transcriptional gene regulation, epigenetic modifications and further processes. TREND-DB visualizes the dynamics of transcriptome 3' end diversification (TREND) in a highly interactive manner; it provides a global APA network map and allows interrogating genes affected by specific APA-regulators and vice versa. It also permits condition-specific functional enrichment analyses of APA-affected genes, which suggest wide biological and clinical relevance across all RNAi conditions. The implementation of the UCSC Genome Browser provides additional customizable layers of gene regulation accounting for individual transcript isoforms (e.g. epigenetics, miRNA-binding sites and RNA-binding proteins). TREND-DB thereby fosters disentangling the role of APA for various biological programs, including potential disease mechanisms, and helps identify their diagnostic and therapeutic potential.
© The Author(s) 2020. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures


Similar articles
-
TRENDseq-A highly multiplexed high throughput RNA 3' end sequencing for mapping alternative polyadenylation.Methods Enzymol. 2021;655:37-72. doi: 10.1016/bs.mie.2021.03.022. Epub 2021 May 26. Methods Enzymol. 2021. PMID: 34183130
-
RNA Binding Protein CELF2 Regulates Signal-Induced Alternative Polyadenylation by Competing with Enhancers of the Polyadenylation Machinery.Cell Rep. 2019 Sep 10;28(11):2795-2806.e3. doi: 10.1016/j.celrep.2019.08.022. Cell Rep. 2019. PMID: 31509743 Free PMC article.
-
scAPAatlas: an atlas of alternative polyadenylation across cell types in human and mouse.Nucleic Acids Res. 2022 Jan 7;50(D1):D356-D364. doi: 10.1093/nar/gkab917. Nucleic Acids Res. 2022. PMID: 34643729 Free PMC article.
-
Emerging Roles of RNA 3'-end Cleavage and Polyadenylation in Pathogenesis, Diagnosis and Therapy of Human Disorders.Biomolecules. 2020 Jun 17;10(6):915. doi: 10.3390/biom10060915. Biomolecules. 2020. PMID: 32560344 Free PMC article. Review.
-
RNA-binding proteins in regulation of alternative cleavage and polyadenylation.Adv Exp Med Biol. 2014;825:97-127. doi: 10.1007/978-1-4939-1221-6_3. Adv Exp Med Biol. 2014. PMID: 25201104 Review.
Cited by
-
Context-specific regulation and function of mRNA alternative polyadenylation.Nat Rev Mol Cell Biol. 2022 Dec;23(12):779-796. doi: 10.1038/s41580-022-00507-5. Epub 2022 Jul 7. Nat Rev Mol Cell Biol. 2022. PMID: 35798852 Free PMC article. Review.
-
Alternative mRNA polyadenylation regulates macrophage hyperactivation via the autophagy pathway.Cell Mol Immunol. 2024 Dec;21(12):1522-1534. doi: 10.1038/s41423-024-01237-8. Epub 2024 Nov 13. Cell Mol Immunol. 2024. PMID: 39537902 Free PMC article.
-
Coordination of alternative splicing and alternative polyadenylation revealed by targeted long read sequencing.Nat Commun. 2023 Sep 7;14(1):5506. doi: 10.1038/s41467-023-41207-8. Nat Commun. 2023. PMID: 37679364 Free PMC article.
-
Alternative polyadenylation quantitative trait methylation mapping in human cancers provides clues into the molecular mechanisms of APA.Am J Hum Genet. 2024 Mar 7;111(3):562-583. doi: 10.1016/j.ajhg.2024.01.010. Epub 2024 Feb 16. Am J Hum Genet. 2024. PMID: 38367620 Free PMC article.
-
The Bidirectional Link Between RNA Cleavage and Polyadenylation and Genome Stability: Recent Insights From a Systematic Screen.Front Genet. 2022 Apr 28;13:854907. doi: 10.3389/fgene.2022.854907. eCollection 2022. Front Genet. 2022. PMID: 35571036 Free PMC article. Review.
References
-
- Carninci P., Klein S.A., Seidel D.J., Pearson B.D., Singer S.F., Michaels P.J., Cox D.I., Seidel D.J., Eskridge R.E., Peterson T.C. et al. .. The transcriptional landscape of the mammalian genome. Science. 2005; 309:1559–1563. - PubMed
-
- Elkon R., Ugalde A.P., Agami R.. Alternative cleavage and polyadenylation: extent, regulation and function. Nat. Rev. Genet. 2013; 14:496–506. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

