Physiologic RNA targets and refined sequence specificity of coronavirus EndoU
- PMID: 32989044
- PMCID: PMC7668261
- DOI: 10.1261/rna.076604.120
Physiologic RNA targets and refined sequence specificity of coronavirus EndoU
Abstract
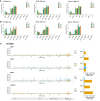
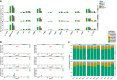
Coronavirus EndoU inhibits dsRNA-activated antiviral responses; however, the physiologic RNA substrates of EndoU are unknown. In this study, we used mouse hepatitis virus (MHV)-infected bone marrow-derived macrophage (BMM) and cyclic phosphate cDNA sequencing to identify the RNA targets of EndoU. EndoU targeted viral RNA, cleaving the 3' side of pyrimidines with a strong preference for U ↓ A and C ↓ A sequences (endoY ↓ A). EndoU-dependent cleavage was detected in every region of MHV RNA, from the 5' NTR to the 3' NTR, including transcriptional regulatory sequences (TRS). Cleavage at two CA dinucleotides immediately adjacent to the MHV poly(A) tail suggests a mechanism to suppress negative-strand RNA synthesis and the accumulation of viral dsRNA. MHV with EndoU (EndoUmut) or 2'-5' phosphodiesterase (PDEmut) mutations provoked the activation of RNase L in BMM, with corresponding cleavage of RNAs by RNase L. The physiologic targets of EndoU are viral RNA templates required for negative-strand RNA synthesis and dsRNA accumulation. Coronavirus EndoU cleaves U ↓ A and C ↓ A sequences (endoY ↓ A) within viral (+) strand RNA to evade dsRNA-activated host responses.
Keywords: coronavirus; dsRNA; endoribonuclease; innate immunity; mouse hepatitis virus.
© 2020 Ancar et al.; Published by Cold Spring Harbor Laboratory Press for the RNA Society.
Figures






References
-
- Alexa A, Rahnenfuhrer J. 2019. topGO: Enrichment analysis for gene ontology (R package version 2.37.0). Bioconductor, https://www.bioconductor.org
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
