Development of a Pseudomonas putida cell-free protein synthesis platform for rapid screening of gene regulatory elements
- PMID: 32995512
- PMCID: PMC7445763
- DOI: 10.1093/synbio/ysy003
Development of a Pseudomonas putida cell-free protein synthesis platform for rapid screening of gene regulatory elements
Abstract
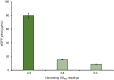
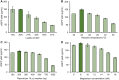
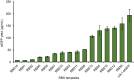
Cell-free protein synthesis (CFPS) systems enable the production of protein without the use of living, intact cells. An emerging area of interest is to use CFPS systems to characterize individual elements for genetic programs [e.g. promoters, ribosome binding sites (RBS)]. To enable this research area, robust CFPS systems must be developed from new chassis organisms. One such chassis is the Gram-negative Pseudomonas bacteria, which have been studied extensively for their diverse metabolism with promises in the field of bioremediation and biosynthesis. Here, we report the development and optimization of a high-yielding (198 ± 5.9 µg/ml) batch CFPS system from Pseudomonas putida ATCC 12633. Importantly, both circular and linear DNA templates can be applied directly to the CFPS reaction to program protein synthesis. Therefore, it is possible to prepare hundreds or even thousands of DNA templates without time-consuming cloning work. This opens the possibility to rapidly assess and validate genetic part performance in vitro before performing experiments in cells. To validate the P. putida CFPS system as a platform for prototyping genetic parts, we designed and constructed a library consisting of 15 different RBSs upstream of the reporter protein sfGFP, which covered an order of magnitude range in expression. Looking forward, our P. putida CFPS platform will not only expand the protein synthesis toolkit for synthetic biology but also serve as a platform in expediting the screening and prototyping of gene regulatory elements.
Keywords: Pseudomonas putida; TX-TL; cell-free protein synthesis; gene regulatory elements; prototyping; synthetic biology.
© The Author(s) 2018. Published by Oxford University Press.
Figures

Similar articles
-
Establishing a Eukaryotic Pichia pastoris Cell-Free Protein Synthesis System.Front Bioeng Biotechnol. 2020 Jun 18;8:536. doi: 10.3389/fbioe.2020.00536. eCollection 2020. Front Bioeng Biotechnol. 2020. PMID: 32626695 Free PMC article.
-
Establishing a High-Yielding Cell-Free Protein Synthesis Platform Derived from Vibrio natriegens.ACS Synth Biol. 2018 Sep 21;7(9):2245-2255. doi: 10.1021/acssynbio.8b00252. Epub 2018 Sep 6. ACS Synth Biol. 2018. PMID: 30107122
-
Establishing a High-Yield Bacillus subtilis-Based Cell-Free Protein Synthesis System for In Vitro Prototyping and Natural Product Biosynthesis.ACS Synth Biol. 2025 Apr 18;14(4):1288-1297. doi: 10.1021/acssynbio.5c00021. Epub 2025 Apr 9. ACS Synth Biol. 2025. PMID: 40203238
-
Cell-Free Synthetic Biology: Engineering Beyond the Cell.Cold Spring Harb Perspect Biol. 2016 Dec 1;8(12):a023853. doi: 10.1101/cshperspect.a023853. Cold Spring Harb Perspect Biol. 2016. PMID: 27742731 Free PMC article. Review.
-
A User's Guide to Cell-Free Protein Synthesis.Methods Protoc. 2019 Mar 12;2(1):24. doi: 10.3390/mps2010024. Methods Protoc. 2019. PMID: 31164605 Free PMC article. Review.
Cited by
-
Multiplex transcriptional characterizations across diverse bacterial species using cell-free systems.Mol Syst Biol. 2019 Aug;15(8):e8875. doi: 10.15252/msb.20198875. Mol Syst Biol. 2019. PMID: 31464371 Free PMC article.
-
Cell-Free Gene Expression: Methods and Applications.Chem Rev. 2025 Jan 8;125(1):91-149. doi: 10.1021/acs.chemrev.4c00116. Epub 2024 Dec 19. Chem Rev. 2025. PMID: 39700225 Free PMC article. Review.
-
Establishing a Eukaryotic Pichia pastoris Cell-Free Protein Synthesis System.Front Bioeng Biotechnol. 2020 Jun 18;8:536. doi: 10.3389/fbioe.2020.00536. eCollection 2020. Front Bioeng Biotechnol. 2020. PMID: 32626695 Free PMC article.
-
Cell-Free Protein Synthesis: A Promising Option for Future Drug Development.BioDrugs. 2020 Jun;34(3):327-348. doi: 10.1007/s40259-020-00417-y. BioDrugs. 2020. PMID: 32198631 Free PMC article. Review.
-
Cell-free prototyping strategies for enhancing the sustainable production of polyhydroxyalkanoates bioplastics.Synth Biol (Oxf). 2018 Sep 4;3(1):ysy016. doi: 10.1093/synbio/ysy016. eCollection 2018. Synth Biol (Oxf). 2018. PMID: 32995523 Free PMC article.
References
-
- Yang J., Kanter G., Voloshin A., Michel-Reydellet N., Velkeen H., Levy R., Swartz J.R. (2005) Rapid expression of vaccine proteins for B-cell lymphoma in a cell-free system. Biotechnol. Bioeng., 89, 503–511. - PubMed
-
- Min S.E., Lee K.-H., Park S.-W., Yoo T.H., Oh C.H., Park J.-H., Yang S.Y., Kim Y.-S., Kim D.-M. (2016) Cell-free production and streamlined assay of cytosol-penetrating antibodies. Biotechnol. Bioeng., 113, 2107–2112. - PubMed
-
- Stech M., Kubick S. (2015) Cell-free synthesis meets antibody production: a review. Antibodies, 4, 12–33.
-
- Bundy B.C., Franciszkowicz M.J., Swartz J.R. (2008) Escherichia coli-based cell-free synthesis of virus-like particles. Biotechnol. Bioeng., 100, 28–37. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
