Transcriptional landscape of the embryonic chicken Müllerian duct
- PMID: 33008304
- PMCID: PMC7532620
- DOI: 10.1186/s12864-020-07106-8
Transcriptional landscape of the embryonic chicken Müllerian duct
Abstract
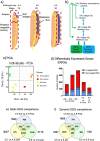
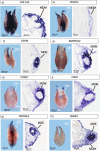
Background: Müllerian ducts are paired embryonic tubes that give rise to the female reproductive tract in vertebrates. Many disorders of female reproduction can be attributed to anomalies of Müllerian duct development. However, the molecular genetics of Müllerian duct formation is poorly understood and most disorders of duct development have unknown etiology. In this study, we describe for the first time the transcriptional landscape of the embryonic Müllerian duct, using the chicken embryo as a model system. RNA sequencing was conducted at 1 day intervals during duct formation to identify developmentally-regulated genes, validated by in situ hybridization.
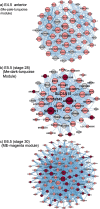
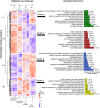
Results: This analysis detected hundreds of genes specifically up-regulated during duct morphogenesis. Gene ontology and pathway analysis revealed enrichment for developmental pathways associated with cell adhesion, cell migration and proliferation, ERK and WNT signaling, and, interestingly, axonal guidance. The latter included factors linked to neuronal cell migration or axonal outgrowth, such as Ephrin B2, netrin receptor, SLIT1 and class A semaphorins. A number of transcriptional modules were identified that centred around key hub genes specifying matrix-associated signaling factors; SPOCK1, HTRA3 and ADGRD1. Several novel regulators of the WNT and TFG-β signaling pathway were identified in Müllerian ducts, including APCDD1 and DKK1, BMP3 and TGFBI. A number of novel transcription factors were also identified, including OSR1, FOXE1, PRICKLE1, TSHZ3 and SMARCA2. In addition, over 100 long non-coding RNAs (lncRNAs) were expressed during duct formation.
Conclusions: This study provides a rich resource of new candidate genes for Müllerian duct development and its disorders. It also sheds light on the molecular pathways engaged during tubulogenesis, a fundamental process in embryonic development.
Keywords: Chicken embryo; FOXE1; Müllerian duct; OSR1; RNA-seq; Sex determination.
Conflict of interest statement
The authors declare that they have no competing interests.
Figures




Similar articles
-
Mini review: Asymmetric Müllerian duct development in the chicken embryo.Front Cell Dev Biol. 2024 Feb 5;12:1347711. doi: 10.3389/fcell.2024.1347711. eCollection 2024. Front Cell Dev Biol. 2024. PMID: 38380340 Free PMC article. Review.
-
Adhesion G-protein-coupled receptor, GPR56, is required for Müllerian duct development in the chick.J Endocrinol. 2020 Feb;244(2):395-413. doi: 10.1530/JOE-19-0419. J Endocrinol. 2020. PMID: 31829965
-
DMRT1 is required for Müllerian duct formation in the chicken embryo.Dev Biol. 2015 Apr 15;400(2):224-36. doi: 10.1016/j.ydbio.2015.02.001. Epub 2015 Feb 12. Dev Biol. 2015. PMID: 25684667
-
The cell biology and molecular genetics of Müllerian duct development.Wiley Interdiscip Rev Dev Biol. 2018 May;7(3):e310. doi: 10.1002/wdev.310. Epub 2018 Jan 19. Wiley Interdiscip Rev Dev Biol. 2018. PMID: 29350886 Review.
-
Identification of differentially expressed genes involved in the regression and development of the chicken Müllerian duct.Int J Dev Biol. 2008;52(8):1135-41. doi: 10.1387/ijdb.072441yh. Int J Dev Biol. 2008. PMID: 18956347
Cited by
-
Mini review: Asymmetric Müllerian duct development in the chicken embryo.Front Cell Dev Biol. 2024 Feb 5;12:1347711. doi: 10.3389/fcell.2024.1347711. eCollection 2024. Front Cell Dev Biol. 2024. PMID: 38380340 Free PMC article. Review.
-
Single-cell transcriptome reveals insights into the development and function of the zebrafish ovary.Elife. 2022 May 19;11:e76014. doi: 10.7554/eLife.76014. Elife. 2022. PMID: 35588359 Free PMC article.
-
Gene expression of male pathway genes sox9 and amh during early sex differentiation in a reptile departs from the classical amniote model.BMC Genomics. 2023 May 5;24(1):243. doi: 10.1186/s12864-023-09334-0. BMC Genomics. 2023. PMID: 37147622 Free PMC article.
-
Overview of Avian Sex Reversal.Int J Mol Sci. 2023 May 5;24(9):8284. doi: 10.3390/ijms24098284. Int J Mol Sci. 2023. PMID: 37175998 Free PMC article. Review.
-
Unveiling Comparative Genomic Trajectories of Selection and Key Candidate Genes in Egg-Type Russian White and Meat-Type White Cornish Chickens.Biology (Basel). 2021 Sep 6;10(9):876. doi: 10.3390/biology10090876. Biology (Basel). 2021. PMID: 34571753 Free PMC article.
References
-
- George FW, Wilson JD. Sex determination and differentiation. New York: Raven Press; 1994.
-
- Josso N, Cate RL, Picard JY, Vigier B, di Clemente N, Wilson C, Imbeaud S, Pepinsky RB, Guerrier D, Boussin L, et al. Anti-mullerian hormone: the Jost factor. Recent Prog Horm Res. 1993;48:1–59. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

