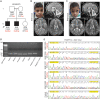
Deficiencies in vesicular transport mediated by TRAPPC4 are associated with severe syndromic intellectual disability.
Van Bergen NJ, Guo Y, Al-Deri N, Lipatova Z, Stanga D, Zhao S, Murtazina R, Gyurkovska V, Pehlivan D, Mitani T, Gezdirici A, Antony J, Collins F, Willis MJH, Coban Akdemir ZH, Liu P, Punetha J, Hunter JV, Jhangiani SN, Fatih JM, Rosenfeld JA, Posey JE, Gibbs RA, Karaca E, Massey S, Ranasinghe TG, Sleiman P, Troedson C, Lupski JR, Sacher M, Segev N, Hakonarson H, Christodoulou J.
Van Bergen NJ, et al.
Brain. 2020 Jan 1;143(1):112-130. doi: 10.1093/brain/awz374.
Brain. 2020.
PMID: 31794024
Free PMC article.