Ancient RNA virus epidemics through the lens of recent adaptation in human genomes
- PMID: 33012231
- PMCID: PMC7702803
- DOI: 10.1098/rstb.2019.0575
Ancient RNA virus epidemics through the lens of recent adaptation in human genomes
Abstract
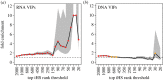
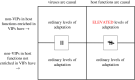
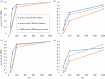
Over the course of the last several million years of evolution, humans probably have been plagued by hundreds or perhaps thousands of epidemics. Little is known about such ancient epidemics and a deep evolutionary perspective on current pathogenic threats is lacking. The study of past epidemics has typically been limited in temporal scope to recorded history, and in physical scope to pathogens that left sufficient DNA behind, such as Yersinia pestis during the Great Plague. Host genomes, however, offer an indirect way to detect ancient epidemics beyond the current temporal and physical limits. Arms races with pathogens have shaped the genomes of the hosts by driving a large number of adaptations at many genes, and these signals can be used to detect and further characterize ancient epidemics. Here, we detect the genomic footprints left by ancient viral epidemics that took place in the past approximately 50 000 years in the 26 human populations represented in the 1000 Genomes Project. By using the enrichment in signals of adaptation at approximately 4500 host loci that interact with specific types of viruses, we provide evidence that RNA viruses have driven a particularly large number of adaptive events across diverse human populations. These results suggest that different types of viruses may have exerted different selective pressures during human evolution. Knowledge of these past selective pressures will provide a deeper evolutionary perspective on current pathogenic threats. This article is part of the theme issue 'Insights into health and disease from ancient biomolecules'.
Keywords: ancient epidemics; genomic adaptation; human evolution.
Conflict of interest statement
We declare we have no competing interests.
Figures

References
Publication types
MeSH terms
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous
