Neural Network Deconvolution Method for Resolving Pathway-Level Progression of Tumor Clonal Expression Programs With Application to Breast Cancer Brain Metastases
- PMID: 33013452
- PMCID: PMC7499245
- DOI: 10.3389/fphys.2020.01055
Neural Network Deconvolution Method for Resolving Pathway-Level Progression of Tumor Clonal Expression Programs With Application to Breast Cancer Brain Metastases
Abstract
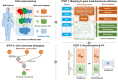
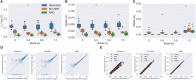
Metastasis is the primary mechanism by which cancer results in mortality and there are currently no reliable treatment options once it occurs, making the metastatic process a critical target for new diagnostics and therapeutics. Treating metastasis before it appears is challenging, however, in part because metastases may be quite distinct genomically from the primary tumors from which they presumably emerged. Phylogenetic studies of cancer development have suggested that changes in tumor genomics over stages of progression often result from shifts in the abundance of clonal cellular populations, as late stages of progression may derive from or select for clonal populations rare in the primary tumor. The present study develops computational methods to infer clonal heterogeneity and dynamics across progression stages via deconvolution and clonal phylogeny reconstruction of pathway-level expression signatures in order to reconstruct how these processes might influence average changes in genomic signatures over progression. We show, via application to a study of gene expression in a collection of matched breast primary tumor and metastatic samples, that the method can infer coarse-grained substructure and stromal infiltration across the metastatic transition. The results suggest that genomic changes observed in metastasis, such as gain of the ErbB signaling pathway, are likely caused by early events in clonal evolution followed by expansion of minor clonal populations in metastasis, a finding that may have translational implications for early detection or prevention of metastasis.
Keywords: brain metastases; breast cancer; deconvolution; gene modules; matrix factorization; pathways; phylogenetics; transcriptome.
Copyright © 2020 Tao, Lei, Lee, Ma and Schwartz.
Figures


Similar articles
-
Deconvolution and phylogeny inference of structural variations in tumor genomic samples.Bioinformatics. 2018 Jul 1;34(13):i357-i365. doi: 10.1093/bioinformatics/bty270. Bioinformatics. 2018. PMID: 29950001 Free PMC article.
-
Robust and accurate deconvolution of tumor populations uncovers evolutionary mechanisms of breast cancer metastasis.Bioinformatics. 2020 Jul 1;36(Suppl_1):i407-i416. doi: 10.1093/bioinformatics/btaa396. Bioinformatics. 2020. PMID: 32657393 Free PMC article.
-
Spatio-Temporal Genomic Heterogeneity, Phylogeny, and Metastatic Evolution in Salivary Adenoid Cystic Carcinoma.J Natl Cancer Inst. 2017 Oct 1;109(10):djx033. doi: 10.1093/jnci/djx033. J Natl Cancer Inst. 2017. PMID: 29117356 Free PMC article.
-
Molecular Profiles of Brain Metastases: A Focus on Heterogeneity.Cancers (Basel). 2021 May 28;13(11):2645. doi: 10.3390/cancers13112645. Cancers (Basel). 2021. PMID: 34071176 Free PMC article. Review.
-
Heterogeneity and tumor evolution reflected in liquid biopsy in metastatic breast cancer patients: a review.Cancer Metastasis Rev. 2022 Jun;41(2):433-446. doi: 10.1007/s10555-022-10023-9. Epub 2022 Mar 14. Cancer Metastasis Rev. 2022. PMID: 35286542 Review.
Cited by
-
Assessing the contribution of tumor mutational phenotypes to cancer progression risk.PLoS Comput Biol. 2021 Mar 12;17(3):e1008777. doi: 10.1371/journal.pcbi.1008777. eCollection 2021 Mar. PLoS Comput Biol. 2021. PMID: 33711014 Free PMC article.
-
Semi-deconvolution of bulk and single-cell RNA-seq data with application to metastatic progression in breast cancer.Bioinformatics. 2022 Jun 24;38(Suppl 1):i386-i394. doi: 10.1093/bioinformatics/btac262. Bioinformatics. 2022. PMID: 35758822 Free PMC article.
-
COVID-19 deep classification network based on convolution and deconvolution local enhancement.Comput Biol Med. 2021 Aug;135:104588. doi: 10.1016/j.compbiomed.2021.104588. Epub 2021 Jun 22. Comput Biol Med. 2021. PMID: 34182330 Free PMC article.
References
-
- Amaratunga D., Cabrera J. (2001). Analysis of data from viral DNA microchips. J. Am. Stat. Assoc. 96, 1161–1170. 10.1198/016214501753381814 - DOI
-
- Bell R. M., Koren Y. (2007). Scalable collaborative filtering with jointly derived neighborhood interpolation weights, in Seventh IEEE International Conference on Data Mining (ICDM 2007) (Omaha, NE: ), 43–52. 10.1109/ICDM.2007.90 - DOI
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

