Iron Regulatory Mechanisms in Saccharomyces cerevisiae
- PMID: 33013818
- PMCID: PMC7509046
- DOI: 10.3389/fmicb.2020.582830
Iron Regulatory Mechanisms in Saccharomyces cerevisiae
Abstract
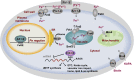
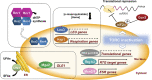
Iron is an essential micronutrient for all eukaryotic organisms because it participates as a redox cofactor in many cellular processes. However, excess iron can damage cells since it promotes the generation of reactive oxygen species. The budding yeast Saccharomyces cerevisiae has been used as a model organism to study the adaptation of eukaryotic cells to changes in iron availability. Upon iron deficiency, yeast utilizes two transcription factors, Aft1 and Aft2, to activate the expression of a set of genes known as the iron regulon, which are implicated in iron uptake, recycling and mobilization. Moreover, Aft1 and Aft2 activate the expression of Cth2, an mRNA-binding protein that limits the expression of genes encoding for iron-containing proteins or that participate in iron-using processes. Cth2 contributes to prioritize iron utilization in particular pathways over other highly iron-consuming and non-essential processes including mitochondrial respiration. Recent studies have revealed that iron deficiency also alters many other metabolic routes including amino acid and lipid synthesis, the mitochondrial retrograde response, transcription, translation and deoxyribonucleotide synthesis; and activates the DNA damage and general stress responses. At high iron levels, the yeast Yap5, Msn2, and Msn4 transcription factors activate the expression of a vacuolar iron importer called Ccc1, which is the most important high-iron protecting factor devoted to detoxify excess cytosolic iron that is stored into the vacuole for its mobilization upon scarcity. The complete sequencing and annotation of many yeast genomes is starting to unveil the diversity and evolution of the iron homeostasis network in this species.
Keywords: Saccharomyces cerevisiae; iron deficiency; iron excess; iron homeostasis; iron metabolism; post-transcriptional regulation; transcriptional regulation; yeast.
Copyright © 2020 Ramos-Alonso, Romero, Martínez-Pastor and Puig.
Figures
Similar articles
-
Iron toxicity in yeast: transcriptional regulation of the vacuolar iron importer Ccc1.Curr Genet. 2018 Apr;64(2):413-416. doi: 10.1007/s00294-017-0767-7. Epub 2017 Oct 17. Curr Genet. 2018. PMID: 29043483 Free PMC article. Review.
-
The glucose sensor Snf1 and the transcription factors Msn2 and Msn4 regulate transcription of the vacuolar iron importer gene CCC1 and iron resistance in yeast.J Biol Chem. 2017 Sep 15;292(37):15577-15586. doi: 10.1074/jbc.M117.802504. Epub 2017 Jul 31. J Biol Chem. 2017. PMID: 28760824 Free PMC article.
-
Phosphorylation and Proteasome Recognition of the mRNA-Binding Protein Cth2 Facilitates Yeast Adaptation to Iron Deficiency.mBio. 2018 Sep 18;9(5):e01694-18. doi: 10.1128/mBio.01694-18. mBio. 2018. PMID: 30228242 Free PMC article.
-
Structure and function of the vacuolar Ccc1/VIT1 family of iron transporters and its regulation in fungi.Comput Struct Biotechnol J. 2020 Nov 23;18:3712-3722. doi: 10.1016/j.csbj.2020.10.044. eCollection 2020. Comput Struct Biotechnol J. 2020. PMID: 33304466 Free PMC article. Review.
-
Mechanisms of iron sensing and regulation in the yeast Saccharomyces cerevisiae.World J Microbiol Biotechnol. 2017 Apr;33(4):75. doi: 10.1007/s11274-017-2215-8. Epub 2017 Mar 17. World J Microbiol Biotechnol. 2017. PMID: 28315258 Review.
Cited by
-
Identification of a divalent metal transporter required for cellular iron metabolism in malaria parasites.Proc Natl Acad Sci U S A. 2024 Nov 5;121(45):e2411631121. doi: 10.1073/pnas.2411631121. Epub 2024 Oct 28. Proc Natl Acad Sci U S A. 2024. PMID: 39467134 Free PMC article.
-
Dietary change without caloric restriction maintains a youthful profile in ageing yeast.PLoS Biol. 2023 Aug 29;21(8):e3002245. doi: 10.1371/journal.pbio.3002245. eCollection 2023 Aug. PLoS Biol. 2023. PMID: 37643155 Free PMC article.
-
Identification of a divalent metal transporter required for cellular iron metabolism in malaria parasites.bioRxiv [Preprint]. 2024 Jun 10:2024.05.10.587216. doi: 10.1101/2024.05.10.587216. bioRxiv. 2024. Update in: Proc Natl Acad Sci U S A. 2024 Nov 5;121(45):e2411631121. doi: 10.1073/pnas.2411631121. PMID: 38798484 Free PMC article. Updated. Preprint.
-
Characterizing the regulatory effects of H2A.Z and SWR1-C on gene expression during hydroxyurea exposure in Saccharomyces cerevisiae.PLoS Genet. 2025 Jan 21;21(1):e1011566. doi: 10.1371/journal.pgen.1011566. eCollection 2025 Jan. PLoS Genet. 2025. PMID: 39836664 Free PMC article.
-
A genome-to-proteome atlas charts natural variants controlling proteome diversity and forecasts their fitness effects.bioRxiv [Preprint]. 2024 Oct 22:2024.10.18.619054. doi: 10.1101/2024.10.18.619054. bioRxiv. 2024. PMID: 39484408 Free PMC article. Preprint.
References
-
- Belli G., Molina M. M., Garcia-Martinez J., Perez-Ortin J. E., Herrero E. (2004). Saccharomyces cerevisiae glutaredoxin 5-deficient cells subjected to continuous oxidizing conditions are affected in the expression of specific sets of genes. J. Biol. Chem. 279 12386–12395. 10.1074/jbc.m311879200 - DOI - PubMed
Publication types
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

