Identification of New Therapeutic Targets for Gastric Cancer With Bioinformatics
- PMID: 33014013
- PMCID: PMC7461879
- DOI: 10.3389/fgene.2020.00865
Identification of New Therapeutic Targets for Gastric Cancer With Bioinformatics
Abstract
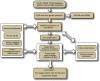
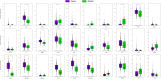
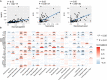
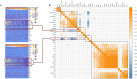
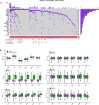
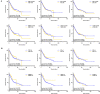
We aimed to identify new targets affecting gastric cancer (GC) prognosis. Six target genes were identified from hub genes based on their relationship with important factors affecting tumor progression, like immune infiltration, purity, tumor mutation burden (TMB), and tumor microenvironment (TME) score. The effect of target genes' somatic mutations and copy number alteration (CNA) was examined to determine their effect on GC prognosis. Six target genes (FBN1, FN1, HGF, MMP9, THBS1, and VCAN) were identified. Reduced expression of each target gene, except MMP9, indicated better prognosis and lower grade in GC. FBN1, THBS1, and VCAN showed lower expression in stage I GC. Non-silencing mutations of the six genes played a role in significantly higher TMB and TME scores. THBS1 mutation was associated with earlier stage GC, and VCAN mutation was associated with lower grade GC. However, patients with target gene CNA displayed higher tumor purity. MMP9, THBS1, and VCAN CNA was associated with lower grade GC, while FBN1 CNA reflected earlier T stage. Additionally, the target genes may affect GC prognosis by influencing multiple oncogenic signaling pathways. FBN1, FN1, HGF, MMP9, THBS1, and VCAN may be new GC prognostic targets by affecting tumor purity, TMB, TME score, and multiple oncogenic signaling pathways.
Keywords: CNA; TMB; TME score; gastric cancer; mutation; prognosis; purity.
Copyright © 2020 Li, Wang, Zhang, Wang and Li.
Figures






Similar articles
-
Identification of cuproptosis-related subtypes, construction of a prognosis model, and tumor microenvironment landscape in gastric cancer.Front Immunol. 2022 Nov 21;13:1056932. doi: 10.3389/fimmu.2022.1056932. eCollection 2022. Front Immunol. 2022. PMID: 36479114 Free PMC article.
-
Bioinformatics analysis of the prognosis and biological significance of VCAN in gastric cancer.Immun Inflamm Dis. 2021 Jun;9(2):547-559. doi: 10.1002/iid3.414. Epub 2021 Feb 25. Immun Inflamm Dis. 2021. PMID: 33631054 Free PMC article.
-
Comprehensive analysis of tumor mutation burden and immune microenvironment in gastric cancer.Biosci Rep. 2021 Feb 26;41(2):BSR20203336. doi: 10.1042/BSR20203336. Biosci Rep. 2021. PMID: 33492335 Free PMC article.
-
Chemokine receptor 4 expression is correlated with the occurrence and prognosis of gastric cancer.FEBS Open Bio. 2020 Jun;10(6):1149-1161. doi: 10.1002/2211-5463.12864. Epub 2020 May 7. FEBS Open Bio. 2020. PMID: 32306562 Free PMC article.
-
Mutation of MUC16 Is Associated With Tumor Mutational Burden and Lymph Node Metastasis in Patients With Gastric Cancer.Front Med (Lausanne). 2022 Feb 8;9:836892. doi: 10.3389/fmed.2022.836892. eCollection 2022. Front Med (Lausanne). 2022. PMID: 35211490 Free PMC article.
Cited by
-
Comprehensive bioinformatic analysis reveals a cancer-associated fibroblast gene signature as a poor prognostic factor and potential therapeutic target in gastric cancer.BMC Cancer. 2022 Jun 23;22(1):692. doi: 10.1186/s12885-022-09736-5. BMC Cancer. 2022. PMID: 35739492 Free PMC article.
-
In Silico Insights Reveal Fibronectin 1 as a Theranostic Marker in Gastric Cancer.Int J Mol Sci. 2024 Oct 16;25(20):11113. doi: 10.3390/ijms252011113. Int J Mol Sci. 2024. PMID: 39456895 Free PMC article.
-
Versican enrichment predicts poor prognosis and response to adjuvant therapy and immunotherapy in gastric cancer.Front Immunol. 2022 Sep 20;13:960570. doi: 10.3389/fimmu.2022.960570. eCollection 2022. Front Immunol. 2022. PMID: 36203562 Free PMC article.
-
A novel stratification framework based on anoikis-related genes for predicting the prognosis in patients with osteosarcoma.Front Immunol. 2023 Jul 27;14:1199869. doi: 10.3389/fimmu.2023.1199869. eCollection 2023. Front Immunol. 2023. PMID: 37575253 Free PMC article.
-
Single-cell RNA-seq and bulk RNA-seq explore the prognostic value of exhausted T cells in hepatocellular carcinoma.IET Syst Biol. 2023 Aug;17(4):228-244. doi: 10.1049/syb2.12072. Epub 2023 Jul 11. IET Syst Biol. 2023. PMID: 37431788 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Miscellaneous

