Peptide Binding to HLA-E Molecules in Humans, Nonhuman Primates, and Mice Reveals Unique Binding Peptides but Remarkably Conserved Anchor Residues
- PMID: 33020145
- PMCID: PMC7653511
- DOI: 10.4049/jimmunol.2000810
Peptide Binding to HLA-E Molecules in Humans, Nonhuman Primates, and Mice Reveals Unique Binding Peptides but Remarkably Conserved Anchor Residues
Abstract
Ag presentation via the nonclassical MHC class Ib molecule HLA-E, with nearly complete identity between the two alleles expressed in humans, HLA-E*01:01 and HLA-E*01:03, can lead to the activation of unconventional T cells in humans. Despite this virtual genetic monomorphism, differences in peptide repertoires binding to the two allelic variants have been reported. To further dissect and compare peptide binding to HLA-E*01:01 and HLA-E*01:03, we used an UV-mediated peptide exchange binding assay and an HPLC-based competition binding assay. In addition, we investigated binding of these same peptides to Mamu-E, the nonhuman primate homologue of human HLA-E, and to the HLA-E-like molecule Qa-1b in mice. We next exploited the differences and homologies in the peptide binding pockets of these four molecules to identify allele specific as well as common features of peptide binding motifs across species. Our results reveal differences in peptide binding preferences and intensities for each human HLA-E variant compared with Mamu-E and Qa-1b Using extended peptide libraries, we identified and refined the peptide binding motifs for each of the four molecules and found that they share main anchor positions, evidenced by conserved amino acid preferences across the four HLA-E molecules studied. In addition, we also identified differences in peptide binding motifs, which could explain the observed variations in peptide binding preferences and affinities for each of the four HLA-E-like molecules. Our results could help with guiding the selection of candidate pathogen-derived peptides with the capacity to target HLA-E-restricted T cells that could be mobilized in vaccination and immunotherapeutic strategies.
Copyright © 2020 by The American Association of Immunologists, Inc.
Conflict of interest statement
The authors have no financial conflicts of interest.
Figures
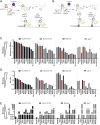



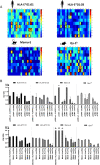

References
-
- Joosten S. A., Ottenhoff T. H. M., Lewinsohn D. M., Hoft D. F., Moody D. B., Seshadri C., Collaboration for Tuberculosis Vaccine Discovery - Donor-Unrestricted T-cells Working Group, Bill and Melinda Gates Foundation 2019. Harnessing donor unrestricted T-cells for new vaccines against tuberculosis. Vaccine 37: 3022–3030. - PMC - PubMed
-
- Geraghty D. E., Stockschleader M., Ishitani A., Hansen J. A. 1992. Polymorphism at the HLA-E locus predates most HLA-A and -B polymorphism. Hum. Immunol. 33: 174–184. - PubMed
-
- Braud V. M., Allan D. S., O’Callaghan C. A., Söderström K., D’Andrea A., Ogg G. S., Lazetic S., Young N. T., Bell J. I., Phillips J. H., et al. 1998. HLA-E binds to natural killer cell receptors CD94/NKG2A, B and C. Nature 391: 795–799. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

