Gene Editing by Extracellular Vesicles
- PMID: 33028045
- PMCID: PMC7582630
- DOI: 10.3390/ijms21197362
Gene Editing by Extracellular Vesicles
Abstract
CRISPR/Cas technologies have advanced dramatically in recent years. Many different systems with new properties have been characterized and a plethora of hybrid CRISPR/Cas systems able to modify the epigenome, regulate transcription, and correct mutations in DNA and RNA have been devised. However, practical application of CRISPR/Cas systems is severely limited by the lack of effective delivery tools. In this review, recent advances in developing vehicles for the delivery of CRISPR/Cas in the form of ribonucleoprotein complexes are outlined. Most importantly, we emphasize the use of extracellular vesicles (EVs) for CRISPR/Cas delivery and describe their unique properties: biocompatibility, safety, capacity for rational design, and ability to cross biological barriers. Available molecular tools that enable loading of desired protein and/or RNA cargo into the vesicles in a controllable manner and shape the surface of EVs for targeted delivery into specific tissues (e.g., using targeting ligands, peptides, or nanobodies) are discussed. Opportunities for both endogenous (intracellular production of CRISPR/Cas) and exogenous (post-production) loading of EVs are presented.
Keywords: biodistribution; exosomes; gene editing; nanoblades, stem cells, mesenchymal stem cells.; nanomedicines; nanoparticles; nanovesicles; pharmacokinetics.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
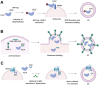

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

