Formicamycin biosynthesis involves a unique reductive ring contraction
- PMID: 33033611
- PMCID: PMC7504897
- DOI: 10.1039/d0sc01712d
Formicamycin biosynthesis involves a unique reductive ring contraction
Abstract
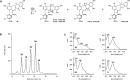
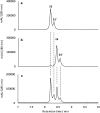
Fasamycin natural products are biosynthetic precursors of the formicamycins. Both groups of compounds are polyketide natural products that exhibit potent antibacterial activity despite displaying different three-dimensional topologies. We show here that transformation of fasamycin into formicamycin metabolites requires two gene products and occurs via a novel two-step ring expansion-ring contraction pathway. Deletion of forX, encoding a flavin dependent monooxygenase, abolished formicamycin production and leads to accumulation of fasamycin E. Deletion of the adjacent gene forY, encoding a flavin dependent oxidoreductase, also abolished formicamycin biosynthesis and led to the accumulation of new lactone metabolites that represent Baeyer-Villiger oxidation products of the fasamycins. These results identify ForX as a Baeyer-Villiger monooxygenase capable of dearomatizing ring C of the fasamycins. Through in vivo cross feeding and biomimetic semi-synthesis experiments we showed that these lactone products represent biosynthetic intermediates that are reduced to formicamycins in a unique reductive ring contraction reaction catalyzed by ForY.
This journal is © The Royal Society of Chemistry 2020.
Figures



References
Grants and funding
LinkOut - more resources
Full Text Sources

