Mu Opioids Induce Biased Signaling at the Full-Length Seven Transmembrane C-Terminal Splice Variants of the mu Opioid Receptor Gene, Oprm1
- PMID: 33033993
- PMCID: PMC8026774
- DOI: 10.1007/s10571-020-00973-5
Mu Opioids Induce Biased Signaling at the Full-Length Seven Transmembrane C-Terminal Splice Variants of the mu Opioid Receptor Gene, Oprm1
Erratum in
-
Correction to: Mu Opioids Induce Biased Signaling at the Full-Length Seven Transmembrane C-Terminal Splice Variants of the mu Opioid Receptor Gene, Oprm1.Cell Mol Neurobiol. 2021 Jul;41(5):1075. doi: 10.1007/s10571-020-00990-4. Cell Mol Neurobiol. 2021. PMID: 33128690 Free PMC article. No abstract available.
Abstract
The biased signaling has been extensively studied in the original mu opioid receptor (MOR-1), particularly through G protein and β-arrestin2 signaling pathways. The concept that the G protein pathway is often linked to the therapeutic effect of the drug, while the β-arrestin pathway is associated to the side effects has been proposed to develop biased analgesic compounds with limited side-effects associated with traditional opiates. The mu opioid receptor gene, OPRM1, undergoes extensive alternative pre-mRNA splicing, generating multiple splice variants or isoforms that are conserved from rodent to human. One type of the Oprm1 splice variants are the full-length 7 transmembrane (7TM) C-terminal splice variants, which have identical receptor structures including entire binding pocket, but contain a different intracellular C-terminal tail resulted from 3' alternative splicing. Increasing evidence suggest that these full-length 7TM C-terminal variants play important roles in mu opioid pharmacology, raising questions regarding biased signaling at these multiple C-terminal variants. In the present study, we investigated the effect of different C-terminal variants on mu agonist-induced G protein coupling, β-arrestin2 recruitment, and ultimately, signaling bias. We found that mu agonists produced marked differences in G protein activation and β-arrestin2 recruitment among various C-terminal variants, leading to biased signaling at various level. Particularly, MOR-1O, an exon 7-associated variant, showed greater β-arrestin2 bias for most mu agonists than MOR-1, an exon 4-associated variant. Biased signaling of G protein-coupled receptors has been defined by evidences that different agonists can produce divergent signaling transduction pathways through a single receptor. Our findings that a single mu agonist can induce differential signaling through multiple 7TM splice variants provide a new perspective on biased signaling at least for Oprm1, which perhaps is important for our understanding of the complex mu opioid actions in vivo where all the 7TM splice variants co-exist.
Keywords: Beta-arrestin; Biased signaling; G-protein; Mu opioid receptor; Splicing.
Conflict of interest statement
YXP is a co-scientific founder of Sparian Biosciences. All other authors declare that there is no conflict of interest.
Figures
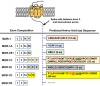
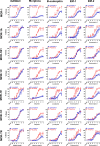

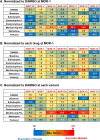
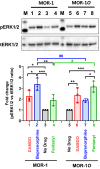
Similar articles
-
Endogenous Opioid Peptides and Alternatively Spliced Mu Opioid Receptor Seven Transmembrane Carboxyl-Terminal Variants.Int J Mol Sci. 2021 Apr 6;22(7):3779. doi: 10.3390/ijms22073779. Int J Mol Sci. 2021. PMID: 33917474 Free PMC article. Review.
-
Identification and characterization of seven new exon 11-associated splice variants of the rat μ opioid receptor gene, OPRM1.Mol Pain. 2011 Jan 21;7:9. doi: 10.1186/1744-8069-7-9. Mol Pain. 2011. PMID: 21255438 Free PMC article.
-
Mediation of buprenorphine analgesia by a combination of traditional and truncated mu opioid receptor splice variants.Synapse. 2016 Oct;70(10):395-407. doi: 10.1002/syn.21914. Epub 2016 Jul 12. Synapse. 2016. PMID: 27223691 Free PMC article.
-
Isolating and characterizing three alternatively spliced mu opioid receptor variants: mMOR-1A, mMOR-1O, and mMOR-1P.Synapse. 2014 Apr;68(4):144-52. doi: 10.1002/syn.21727. Epub 2014 Jan 16. Synapse. 2014. PMID: 24375714 Free PMC article.
-
Alternative Pre-mRNA Splicing of the Mu Opioid Receptor Gene, OPRM1: Insight into Complex Mu Opioid Actions.Biomolecules. 2021 Oct 15;11(10):1525. doi: 10.3390/biom11101525. Biomolecules. 2021. PMID: 34680158 Free PMC article. Review.
Cited by
-
Functional differences in the mu opioid receptor SNP 118A>G are dependent on receptor splice-variant and agonist-specific recruitment of β-arrestin.Clin Transl Sci. 2024 Aug;17(8):e13888. doi: 10.1111/cts.13888. Clin Transl Sci. 2024. PMID: 39118392 Free PMC article.
-
Interactive effects of morphine and the HIV integrase inhibitor, cabotegravir, in male and female mice.Biomed Pharmacother. 2025 Mar;184:117925. doi: 10.1016/j.biopha.2025.117925. Epub 2025 Feb 24. Biomed Pharmacother. 2025. PMID: 39999644 Free PMC article.
-
Exploring Pharmacological Functions of Alternatively Spliced Variants of the Mu Opioid Receptor Gene, Oprm1, via Gene-Targeted Animal Models.Int J Mol Sci. 2022 Mar 10;23(6):3010. doi: 10.3390/ijms23063010. Int J Mol Sci. 2022. PMID: 35328429 Free PMC article. Review.
-
Involvement of the Opioid Peptide Family in Cancer Progression.Biomedicines. 2023 Jul 14;11(7):1993. doi: 10.3390/biomedicines11071993. Biomedicines. 2023. PMID: 37509632 Free PMC article. Review.
-
Endogenous Opioid Peptides and Alternatively Spliced Mu Opioid Receptor Seven Transmembrane Carboxyl-Terminal Variants.Int J Mol Sci. 2021 Apr 6;22(7):3779. doi: 10.3390/ijms22073779. Int J Mol Sci. 2021. PMID: 33917474 Free PMC article. Review.
References
-
- Abbadie C, Pan Y-X, Drake CT, Pasternak GW (2000a) Comparative immunhistochemical distributions of carboxy terminus epitopes from the mu opioid receptor splice variants MOR-1D, MOR-1 and MOR-1C in the mouse and rat central nervous systems. Neuroscience 100:141–153 - PubMed
-
- Abbadie C, Pan Y-X, Pasternak GW (2000b) Differential distribution in rat brain of mu opioid receptor carboxy terminal splice variants MOR-1C and MOR-1-like immunoreactivity: evidence for region-specific processing. J Comp Neurol 419:244–256 - PubMed
-
- Abbadie C, Pasternak GW (2001) Differential in vivo internalization of MOR-1 and MOR-1C by morphine. NeuroReport 12:3069–3072 - PubMed
-
- Barak LS, Ferguson SS, Zhang J, Caron MG (1997) A beta-arrestin/green fluorescent protein biosensor for detecting G protein-coupled receptor activation. J Biol Chem 272:27497–27500. 10.1074/jbc.272.44.27497 - PubMed
-
- Bertrand L et al (2002) The BRET2/arrestin assay in stable recombinant cells: a platform to screen for compounds that interact with G protein-coupled receptors (GPCRS). J Recept Signal Transduct Res 22:533–541. 10.1081/rrs-120014619 - PubMed
MeSH terms
Substances
Grants and funding
- T32 DA007242/DA/NIDA NIH HHS/United States
- DA042888/DA/NIDA NIH HHS/United States
- R01 DA042888/DA/NIDA NIH HHS/United States
- R21 DA046714/DA/NIDA NIH HHS/United States
- P30 CA008748/CA/NCI NIH HHS/United States
- R21 DA029244/DA/NIDA NIH HHS/United States
- CA008748/CA/NCI NIH HHS/United States
- R37 DA007242/DA/NIDA NIH HHS/United States
- DA046714/DA/NIDA NIH HHS/United States
- R01 DA007242/DA/NIDA NIH HHS/United States
- DA007242/DA/NIDA NIH HHS/United States
- R21 DA040858/DA/NIDA NIH HHS/United States
- UG3 DA048379/DA/NIDA NIH HHS/United States
- DA042888/DA/NIDA NIH HHS/United States
- DA046714/DA/NIDA NIH HHS/United States
- DA007242/DA/NIDA NIH HHS/United States
- CA008748/CA/NCI NIH HHS/United States
LinkOut - more resources
Full Text Sources
Research Materials

