Identification of Novel Inhibitors of a Plant Group A Protein Phosphatase Type 2C Using a Combined In Silico and Biochemical Approach
- PMID: 33042170
- PMCID: PMC7524867
- DOI: 10.3389/fpls.2020.526460
Identification of Novel Inhibitors of a Plant Group A Protein Phosphatase Type 2C Using a Combined In Silico and Biochemical Approach
Abstract
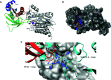
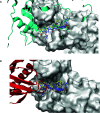
Type 2C protein phosphatases (PP2Cs) of group A play a significant role in the regulation of various processes in plants including growth, development, ion transport, and stress acclimation. In this study, we selected potential PP2C group A inhibitors using a structure-based virtual screening method followed by biochemical and in vitro validation. Over twenty million chemical compounds from the ZINC database were used for docking studies. The precision of the calculations was increased by an induced-fit docking protocol and the molecular mechanics/generalized Born surface area (MM/GBSA) method, which yielded approximate values for the binding energy of the protein-ligand complex. After clustering and ranking their activity, the top-ranking compounds were tested against PP2C group A members in vitro and their in vivo activity was also explored. Phosphatase activity assays identified two compounds with significant inhibitory activity against ABI1 protein ranging from around 57 to 91% at a concentration of 100 μM. Importantly, this in vitro activity correlated well with in vivo inhibition of seed germination, as expected for PP2C inhibitors. The results should promote the design of novel inhibitors with improved potency against ABI1-like and other PP2Cs that might be used in agriculture for the protection of crops against stress.
Keywords: PP2C inhibitor; protein phosphatase 2C group A; protein-ligand complexes; structure-based virtual screening.
Copyright © 2020 Janicki, Marczak, Cieśla and Ludwików.
Figures







References
-
- DeLano W. L. (2002). The PyMOL Molecular Graphics System (San Carlos, CA: DeLanoScientific; ). Available at: www.pymol.org.
LinkOut - more resources
Full Text Sources

